我没有循环的经验,但是看起来我需要创建一些循环来正确分析我的数据。您能否展示如何在我已经创建的代码中创建一个简单的循环?让我们使用循环来获得一些图表:
pdf(file = sprintf("complex I analysis", tbl_comp_abu1), paper='A4r')
ggplot(df_tbl_data1_comp1, aes(Size_Range, Abundance, group=factor(Gene_Name))) +
theme(legend.title=element_blank()) +
geom_line(aes(color=factor(Gene_Name))) +
ggtitle("Data1 - complex I")+
theme(axis.text.x = element_text(angle = 90, hjust = 1))
ggplot(df_tbl_data2_comp1, aes(Size_Range, Abundance, group=factor(Gene_Name))) +
theme(legend.title=element_blank()) +
geom_line(aes(color=factor(Gene_Name))) +
ggtitle("Data2 - complex I")+
theme(axis.text.x = element_text(angle = 90, hjust = 1))
ggplot(df_tbl_data3_comp1, aes(Size_Range, Abundance, group=factor(Gene_Name))) +
theme(legend.title=element_blank()) +
geom_line(aes(color=factor(Gene_Name))) +
ggtitle("Datas3 - complex I")+
theme(axis.text.x = element_text(angle = 90, hjust = 1))
dev.off()
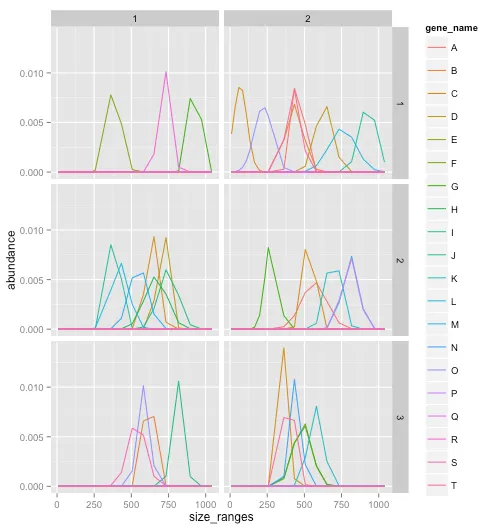
现在的问题是我想要实现什么。首先,我有10个复合物要分析,这意味着需要创建10个pdf文件。示例展示了来自第一个复合物的三个不同数据集的图形。为了正确完成,变量
comp1中的数字(来自df_tbl_dataX_comp1)必须从1更改为10-取决于我们想要绘制哪个复合物。下一个需要通过循环更改的事情是pdf文件的名称和每个图形...是否可能编写这样的循环?数据:
structure(list(Size_Range = structure(c(1L, 1L, 1L, 2L, 2L, 2L,
3L, 3L, 3L, 4L, 4L, 4L, 5L, 5L, 5L, 6L, 6L, 6L, 7L, 7L, 7L, 8L,
8L, 8L, 9L, 9L, 9L, 10L, 10L, 10L, 11L, 11L, 11L, 12L, 12L, 12L,
13L, 13L, 13L, 14L, 14L, 14L, 15L, 15L, 15L, 16L, 16L, 16L, 17L,
17L, 17L, 18L, 18L, 18L, 19L, 19L, 19L, 20L, 20L, 20L), .Label = c("10",
"34", "59", "84", "110", "134", "165", "199", "234", "257", "362",
"433", "506", "581", "652", "733", "818", "896", "972", "1039"
), class = "factor"), Abundance = c(0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 142733.475, 108263.525, 98261.11, 649286.165,
3320759.803, 3708515.148, 6691260.945, 30946562.92, 180974.3725,
4530005.805, 21499827.89, 0, 15032198.54, 4058060.583, 0, 3842964.97,
2544030.857, 0, 1640476.977, 286249.1775, 0, 217388.5675, 1252965.433,
0, 1314666.05, 167467.8825, 0, 253798.15, 107244.9925, 0, 207341.1925,
15755.485, 0, 71015.85, 14828.5075, 0, 25966.2325, 0, 0, 0, 0,
0, 0), Gene_Name = c("AT1G01080", "AT1G01090", "AT1G01320", "AT1G01420",
"AT1G01470", "AT1G01560", "AT1G01800", "AT1G02150", "AT1G02500",
"AT1G02560", "AT1G02780", "AT1G02880", "AT1G02920", "AT1G02930",
"AT1G03030", "AT1G03090", "AT1G03110", "AT1G03130", "AT1G03220",
"AT1G03230", "AT1G03330", "AT1G03475", "AT1G03630", "AT1G03680",
"AT1G03870", "ATCG00420", "ATCG00470", "ATCG00480", "ATCG00490",
"ATCG00500", "ATCG00650", "ATCG00660", "ATCG00670", "ATCG00740",
"ATCG00750", "ATCG00842", "ATCG01100", "ATCG01030", "ATCG01114",
"ATCG01665", "ATCG00770", "ATCG00780", "ATCG00800", "ATCG00810",
"ATCG00820", "ATCG00722", "ATCG00744", "ATCG00855", "ATCG00853",
"ATCG00888", "ATCG00733", "ATCG00766", "ATCG00812", "ATCG00821",
"ATCG00856", "ATCG00830", "ATCG00900", "ATCG01060", "ATCG01110",
"ATCG01120")), .Names = c("Size_Range", "Abundance", "Gene_Name"
), row.names = c(NA, -60L), class = "data.frame")
lapply或类似的东西。 - Herokap = as.list(1:3),p[[1]] = ggplot(...) + ...,p[[2]] = ...然后pdf("plots.pdf", paper = "A4r"); p; dev.off()。 - Akhil Nair