我使用线性判别分析(LDA)来研究一组变量在区分3组方面的效果。然后我使用
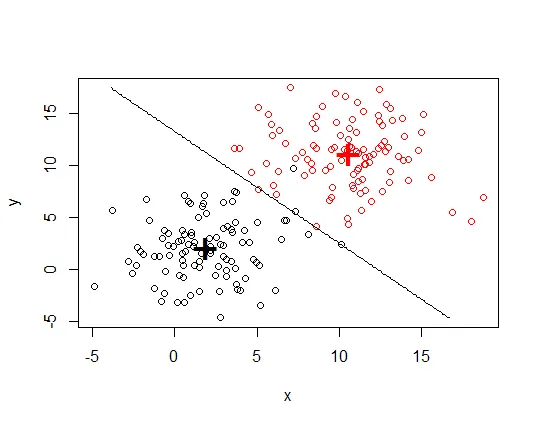
以下是一些示例数据(3组,2个变量):
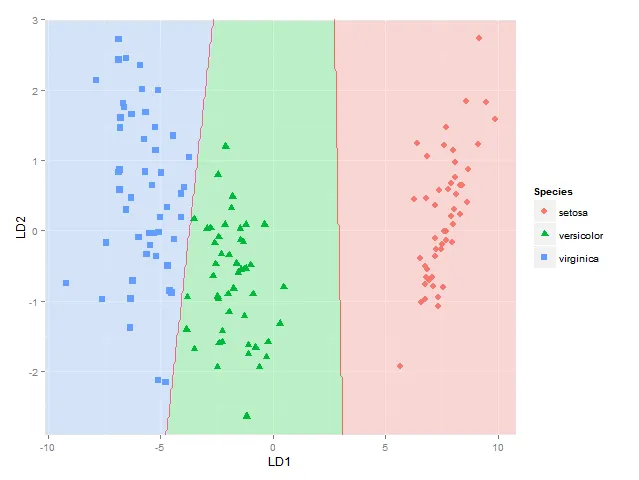
plot.lda() 函数将我的数据绘制在两个线性判别分析图上(LD1 在x轴上,LD2 在y轴上)。现在我想将LDA的分类边界添加到图中,但我没有看到函数中允许这样做的参数。虽然 partimat() 函数允许可视化LD分类边界,但在此情况下,变量用作x和y轴,而不是线性判别分析图。如何向 plot.lda 添加分类边界的任何建议都将不胜感激。以下是一些示例代码:library(MASS)
# LDA
t.lda = lda(Group ~ Var1 + Var2, data=mydata,
na.action="na.omit", CV=TRUE)
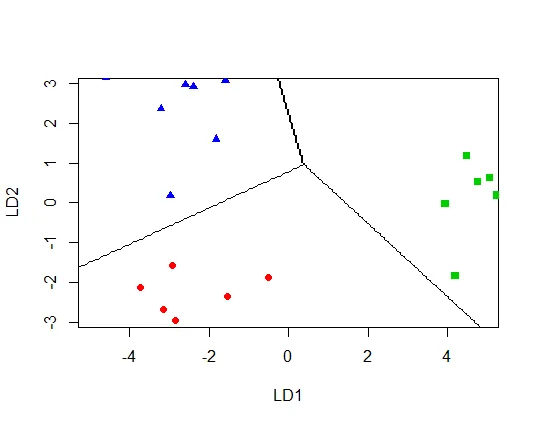
# Scatter plot using the two discriminant dimensions
plot(t.lda,
panel = function(x, y, ...) { points(x, y, ...) },
col = c(4,2,3)[factor(mydata$Group)],
pch = c(17,19,15)[factor(mydata$Group)],
ylim=c(-3,3), xlim=c(-5,5))
以下是一些示例数据(3组,2个变量):
> dput(mydata)
structure(list(Group = c("a", "a", "a", "a", "a", "a", "a", "a",
"b", "b", "b", "b", "b", "b", "b", "b", "c", "c", "c", "c", "c",
"c", "c", "c"), Var1 = c(7.5, 6.9, 6.5, 7.3, 8.1, 8, 7.4, 7.8,
8.3, 8.7, 8.9, 9.3, 8.5, 9.6, 9.8, 9.7, 11.2, 10.9, 11.5, 12,
11, 11.6, 11.7, 11.3), Var2 = c(-6.5, -6.2, -6.7, -6.9, -7.1,
-8, -6.5, -6.3, -9.3, -9.5, -9.6, -9.1, -8.9, -8.7, -9.9, -10,
-6.7, -6.4, -6.8, -6.1, -7.1, -8, -6.9, -6.6)), .Names = c("Group",
"Var1", "Var2"), class = "data.frame", row.names = c(NA, -24L
))
> head(mydata)
Group Var1 Var2
1 a 7.5 -6.5
2 a 6.9 -6.2
3 a 6.5 -6.7
4 a 7.3 -6.9
5 a 8.1 -7.1
6 a 8.0 -8.0
编辑:在Roman的答案之后,我试图修改代码以在线性判别比例尺上绘制分类边界(这是我想要实现的),而不是在原始变量的比例尺上绘制。然而,边界没有停留在它应该停留的地方。如果您对我在这里做错了什么有任何建议,将不胜感激:
#create new data
np = 300
nd.x = seq(from = min(mydata$Var1), to = max(mydata$Var1), length.out = np)
nd.y = seq(from = min(mydata$Var2), to = max(mydata$Var2), length.out = np)
nd = expand.grid(Var1 = nd.x, Var2 = nd.y)
#run lda and predict using new data
new.lda = lda(Group ~ Var1 + Var2, data=mydata)
prd = as.numeric(predict(new.lda, newdata = nd)$class)
#create LD sequences from min - max values
p = predict(new.lda, newdata= nd)
p.x = seq(from = min(p$x[,1]), to = max(p$x[,1]), length.out = np) #LD1 scores
p.y = seq(from = min(p$x[,2]), to = max(p$x[,2]), length.out = np) #LD2 scores
#create original plot
quartz()
plot(t.lda, panel = function(x, y, ...) { points(x, y, ...) },
col = c(4,2,3)[factor(mydata$Group)],
pch = c(17,19,15)[factor(mydata$Group)],
ylim=c(-3,3), xlim=c(-5,5))
#add classification border on scale of linear discriminants (NOTE: this step currently doesn't work)
contour(x = p.x, y = p.y, z = matrix(prd, nrow = np, ncol = np),
levels = c(1, 2, 3), add = TRUE, drawlabels = FALSE)