(我为什么要这样做?请见下面的解释)
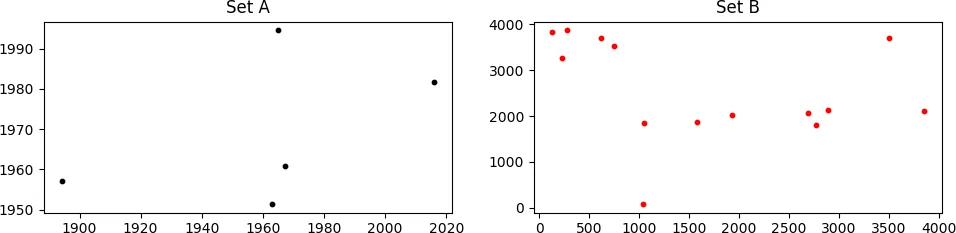
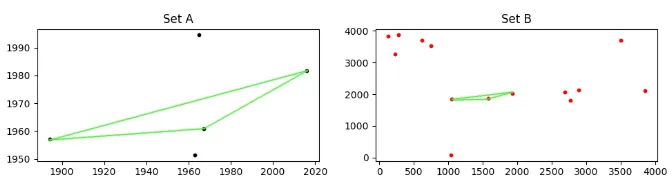
考虑如下所示的两组点,A 和 B
看起来可能不像,但是集合A“隐藏”在集合B中。由于B中的点是相对于A进行缩放、旋转和平移的,因此它们不容易被发现。A中存在的一些点在B中不存在,而B中包含许多不在A中的点,这使得情况更加糟糕。
我需要找到适当的缩放、旋转和平移值,以便将B集合与A集合匹配。在上面的情况中,正确的值为:
scale = 0.14, rot_angle = 0.0, x_transl = 35.0, y_transl = 2.0
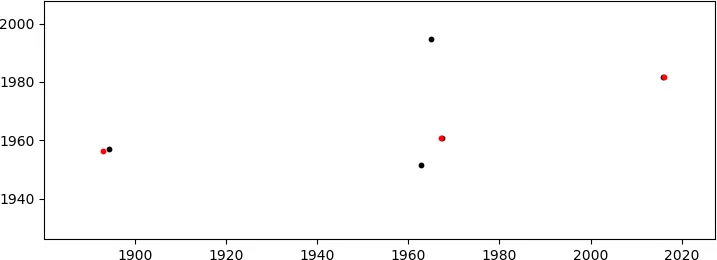
这将产生(足够好的)匹配
(在红色区域中,仅显示匹配的B点;这些点位于右侧第一张图中的扇形区域1000<x<2000,y~2000中)。由于自由度很多(DoF:缩放+旋转+2D平移),我意识到可能存在不匹配的可能性,但是这些点的坐标并不是随机的(尽管看起来像),因此发生这种情况的概率非常小。
我编写的代码(见下文)使用蛮力法循环遍历每个预定义范围中所有可能的DoF值。代码的核心基于将A中的每个点与B中的任何点的距离最小化。
该代码可以工作(实际上生成了上述解决方案),但由于解决方案数量(即每个DoF的接受值的组合)随着范围的增大而增加,因此它可能会变得不可接受地缓慢(还会占用系统中的所有RAM)。
如何提高代码的性能?我可以接受任何解决方案,包括使用numpy和/或scipy。也许可以尝试类似Basing-Hopping这样的方法来搜索最佳匹配(或相对接近的匹配),而不是目前采用的暴力方法?
import numpy as np
from scipy.spatial import distance
import math
def scalePoints(B_center, delta_x, delta_y, scale):
"""
Scales xy points.
http://codereview.stackexchange.com/q/159183/35351
"""
x_scale = B_center[0] - scale * delta_x
y_scale = B_center[1] - scale * delta_y
return x_scale, y_scale
def rotatePoints(center, x, y, angle):
"""
Rotates points in 'xy' around 'center'. Angle is in degrees.
Rotation is counter-clockwise
https://dev59.com/2mIj5IYBdhLWcg3wpmsH#20024348
"""
angle = math.radians(angle)
xy_rot = x - center[0], y - center[1]
xy_rot = (xy_rot[0] * math.cos(angle) - xy_rot[1] * math.sin(angle),
xy_rot[0] * math.sin(angle) + xy_rot[1] * math.cos(angle))
xy_rot = xy_rot[0] + center[0], xy_rot[1] + center[1]
return xy_rot
def distancePoints(set_A, x_transl, y_transl):
"""
Find the sum of the minimum distance of points in set_A to points in set_B.
"""
d = distance.cdist(set_A, zip(*[x_transl, y_transl]), 'euclidean')
# Sum of all minimal distances.
d_sum = np.sum(np.min(d, axis=1))
return d_sum
def match_frames(
set_A, B_center, delta_x, delta_y, tol, sc_min, sc_max, sc_step,
ang_min, ang_max, ang_step, xmin, xmax, xstep, ymin, ymax, ystep):
"""
Process all possible solutions in the defined ranges.
"""
N = len(set_A)
# Ranges
sc_range = np.arange(sc_min, sc_max, sc_step)
ang_range = np.arange(ang_min, ang_max, ang_step)
x_range = np.arange(xmin, xmax, xstep)
y_range = np.arange(ymin, ymax, ystep)
print("Total solutions: {:.2e}".format(
np.prod([len(_) for _ in [sc_range, ang_range, x_range, y_range]])))
d_sum, params_all = [], []
for scale in sc_range:
# Scaled points.
x_scale, y_scale = scalePoints(B_center, delta_x, delta_y, scale)
for ang in ang_range:
# Rotated points.
xy_rot = rotatePoints(B_center, x_scale, y_scale, ang)
# x translation
for x_tr in x_range:
x_transl = xy_rot[0] + x_tr
# y translation
for y_tr in y_range:
y_transl = xy_rot[1] + y_tr
# Find minimum distance sum.
d_sum.append(distancePoints(set_A, x_transl, y_transl))
# Store solutions.
params_all.append([scale, ang, x_tr, y_tr])
# Condition to break out if given tolerance for match
# is achieved.
if d_sum[-1] <= tol * N:
print("Match found:", scale, ang, x_tr, y_tr)
i_min = d_sum.index(min(d_sum))
return i_min, params_all
# Print best solution found so far.
i_min = d_sum.index(min(d_sum))
print("d_sum_min = {:.2f}".format(d_sum[i_min]))
return i_min, params_all
# Data.
set_A = [[2015.81, 1981.665], [1967.31, 1960.865], [1962.91, 1951.365],
[1964.91, 1994.565], [1894.41, 1957.065]]
set_B = [
[2689.28, 3507.04, 2895.67, 1051.3, 1929.49, 1035.97, 752.44, 130.62,
620.06, 2769.06, 1580.77, 281.76, 224.54, 3848.3],
[2061.19, 3700.27, 2131.2, 1837.3, 2017.52, 80.96, 3524.61, 3821.22,
3711.53, 1812.12, 1868.33, 3865.77, 3273.77, 2100.71]]
# This is necessary to apply the scaling.
x, y = np.asarray(set_B)
# Center of B points, defined as the center of the minimal rectangle that
# contains all points.
B_center = [(min(x) + max(x)) * .5, (min(y) + max(y)) * .5]
# Difference between the center coordinates and the xy points.
delta_x, delta_y = B_center[0] - x, B_center[1] - y
# Tolerance in pixels for match.
tol = 1.
# Ranges for each DoF.
sc_min, sc_max, sc_step = .01, .2, .01
ang_min, ang_max, ang_step = -30., 30., 1.
xmin, xmax, xstep = -150., 150., 1.
ymin, ymax, ystep = -150., 150., 1.
# Find proper scaling + rotation + translation for set_B.
i_min, params_all = match_frames(
set_A, B_center, delta_x, delta_y, tol, sc_min, sc_max, sc_step,
ang_min, ang_max, ang_step, xmin, xmax, xstep, ymin, ymax, ystep)
# Best match found
print(params_all[i_min])
我为什么要这样做?
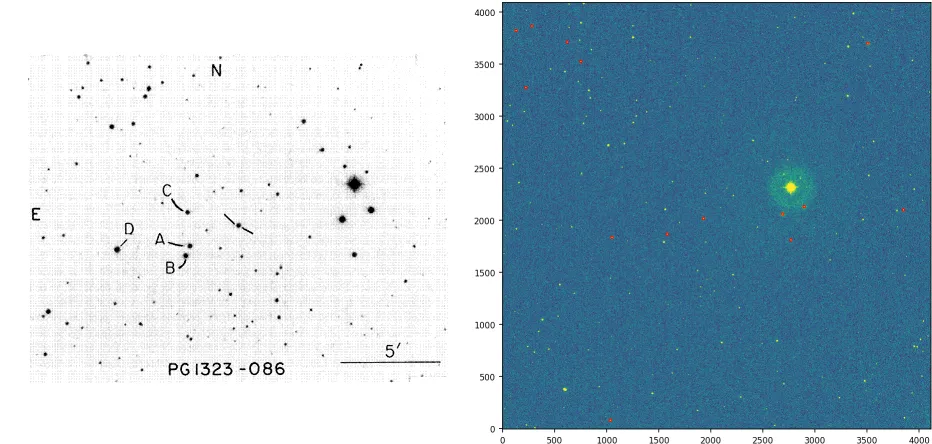
当天文学家观测星场时,还需要观测所谓的“标准星场”。这是为了能够将观测到的星体的“仪器星等”(一种对亮度的对数测量)转换为通用的普遍尺度,因为这些星等取决于所使用的光学系统(望远镜+CCD阵列)。在此示例中,可以看到标准场位于左下方,观测场位于右侧。
请注意,在上面使用的集合A中的点是标准场中标记的星号,而集合B是在观测场中检测到的星星(在上面用红色标记)。将观测场中与标准场中的星星相对应的星星识别过程仍然是通过肉眼完成的,即使是今天也是如此。这是由于任务的复杂性。
在上面的观测图像中,存在相当多的缩放,但没有旋转和很少的平移。这构成了一个相当有利的情况;它可能会更糟。我正在尝试开发一个简单的算法,以避免逐个手动识别观测场中的星星作为标准场中的星星。
litepresence提出的解决方案
这是我根据litepresence的答案编写的脚本。
import itertools
import numpy as np
import matplotlib.pyplot as plt
def getTriangles(set_X, X_combs):
"""
Inefficient way of obtaining the lengths of each triangle's side.
Normalized so that the minimum length is 1.
"""
triang = []
for p0, p1, p2 in X_combs:
d1 = np.sqrt((set_X[p0][0] - set_X[p1][0]) ** 2 +
(set_X[p0][1] - set_X[p1][1]) ** 2)
d2 = np.sqrt((set_X[p0][0] - set_X[p2][0]) ** 2 +
(set_X[p0][1] - set_X[p2][1]) ** 2)
d3 = np.sqrt((set_X[p1][0] - set_X[p2][0]) ** 2 +
(set_X[p1][1] - set_X[p2][1]) ** 2)
d_min = min(d1, d2, d3)
d_unsort = [d1 / d_min, d2 / d_min, d3 / d_min]
triang.append(sorted(d_unsort))
return triang
def sumTriangles(A_triang, B_triang):
"""
For each normalized triangle in A, compare with each normalized triangle
in B. find the differences between their sides, sum their absolute values,
and select the two triangles with the smallest sum of absolute differences.
"""
tr_sum, tr_idx = [], []
for i, A_tr in enumerate(A_triang):
for j, B_tr in enumerate(B_triang):
# Absolute value of lengths differences.
tr_diff = abs(np.array(A_tr) - np.array(B_tr))
# Sum the differences
tr_sum.append(sum(tr_diff))
tr_idx.append([i, j])
# Index of the triangles in A and B with the smallest sum of absolute
# length differences.
tr_idx_min = tr_idx[tr_sum.index(min(tr_sum))]
A_idx, B_idx = tr_idx_min[0], tr_idx_min[1]
print("Smallest difference: {}".format(min(tr_sum)))
return A_idx, B_idx
# Data.
set_A = [[2015.81, 1981.665], [1967.31, 1960.865], [1962.91, 1951.365],
[1964.91, 1994.565], [1894.41, 1957.065]]
set_B = [
[2689.28, 3507.04, 2895.67, 1051.3, 1929.49, 1035.97, 752.44, 130.62,
620.06, 2769.06, 1580.77, 281.76, 224.54, 3848.3],
[2061.19, 3700.27, 2131.2, 1837.3, 2017.52, 80.96, 3524.61, 3821.22,
3711.53, 1812.12, 1868.33, 3865.77, 3273.77, 2100.71]]
set_B = zip(*set_B)
# All possible triangles.
A_combs = list(itertools.combinations(range(len(set_A)), 3))
B_combs = list(itertools.combinations(range(len(set_B)), 3))
# Obtain normalized triangles.
A_triang, B_triang = getTriangles(set_A, A_combs), getTriangles(set_B, B_combs)
# Index of the A and B triangles with the smallest difference.
A_idx, B_idx = sumTriangles(A_triang, B_triang)
# Indexes of points in A and B of the best match triangles.
A_idx_pts, B_idx_pts = A_combs[A_idx], B_combs[B_idx]
print 'triangle A %s matches triangle B %s' % (A_idx_pts, B_idx_pts)
# Matched points in A and B.
print "A:", [set_A[_] for _ in A_idx_pts]
print "B:", [set_B[_] for _ in B_idx_pts]
# Plot
A_pts = zip(*[set_A[_] for _ in A_idx_pts])
B_pts = zip(*[set_B[_] for _ in B_idx_pts])
plt.scatter(*A_pts, s=10, c='k')
plt.scatter(*B_pts, s=10, c='r')
plt.show()
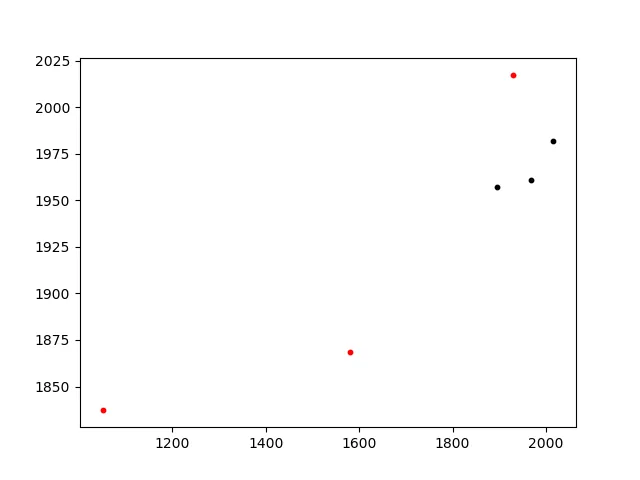
这个方法几乎是即时的,能够产生正确的匹配:
Smallest difference: 0.0314154749597
triangle A (0, 1, 4) matches triangle B (3, 4, 10)
A: [[2015.81, 1981.665], [1967.31, 1960.865], [1894.41, 1957.065]]
B: [(1051.3, 1837.3), (1929.49, 2017.52), (1580.77, 1868.33)]
A到B的点之间的距离)是一个合理的方法,但确实,通过预定义的三个 DOF 值进行暴力迭代并不太好。也许可以看一下 适当的 优化库?http://esa.github.io/pygmo/quickstart.html - Aleksander Lidtkepip、conda或最多git clone安装的软件包。此外,该软件包似乎利用了多个CPU,而我正在寻求代码级别的性能提升。 - Gabriel