我使用 flexmix 进行以下方式的预测:
pred = predict(mod, NPreg)
clust = clusters(mod,NPreg)
result = cbind(NPreg,data.frame(pred),data.frame(clust))
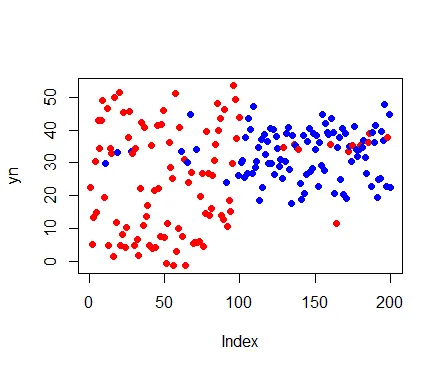
plot(result$yn,col = c("red","blue")[result$clust],pch = 16,ylab = "yn")
混淆矩阵如下:
table(result$class,result$clust)
为了获得yn的预测值,我选择数据点所属聚类的组成值。
for(i in 1:nrow(result)){
result$pred_model1[i] = result[,paste0("Comp.",result$clust[i],".1")][i]
result$pred_model2[i] = result[,paste0("Comp.",result$clust[i],".2")][i]
}
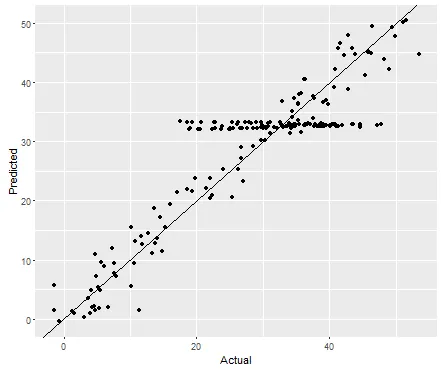
实际结果与预测结果显示拟合度良好(这里只添加一个,因为你的两个模型是相同的,你将使用
pred_model2来表示第二个模型)。
qplot(result$yn, result$pred_model1,xlab="Actual",ylab="Predicted") + geom_abline()
RMSE = sqrt(mean((result$yn-result$pred_model1)^2))
给出了均方根误差为5.54。
这个答案基于我在使用flexmix时阅读的许多SO答案。它对我的问题很有效。
您可能还有兴趣将两个分布可视化。我的模型如下,显示了一些重叠,因为组件比例不接近1。
Call:
flexmix(formula = yn ~ x, data = NPreg, k = 2,
model = list(FLXMRglm(yn ~ x, family = "gaussian"),
FLXMRglm(yn ~ x, family = "gaussian")))
prior size post>0 ratio
Comp.1 0.481 102 129 0.791
Comp.2 0.519 98 171 0.573
'log Lik.' -1312.127 (df=13)
AIC: 2650.255 BIC: 2693.133
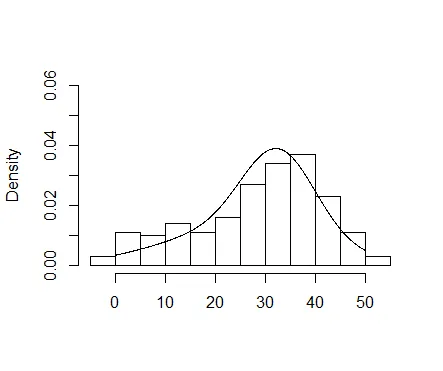
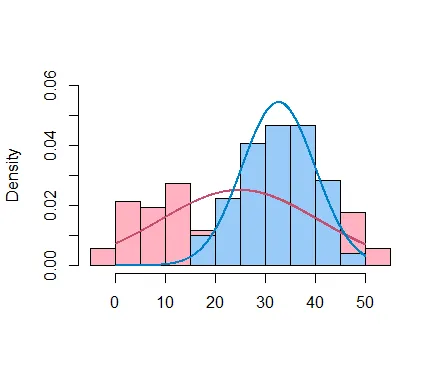
我还使用直方图生成密度分布来可视化两个组件。这得益于betareg维护者在SO上的一个答案的启发。
a = subset(result, clust == 1)
b = subset(result, clust == 2)
hist(a$yn, col = hcl(0, 50, 80), main = "",xlab = "", freq = FALSE, ylim = c(0,0.06))
hist(b$yn, col = hcl(240, 50, 80), add = TRUE,main = "", xlab = "", freq = FALSE, ylim = c(0,0.06))
ys = seq(0, 50, by = 0.1)
lines(ys, dnorm(ys, mean = mean(a$yn), sd = sd(a$yn)), col = hcl(0, 80, 50), lwd = 2)
lines(ys, dnorm(ys, mean = mean(b$yn), sd = sd(b$yn)), col = hcl(240, 80, 50), lwd = 2)
p <- prior(mod)
hist(result$yn, freq = FALSE,main = "", xlab = "",ylim = c(0,0.06))
lines(ys, p[1] * dnorm(ys, mean = mean(a$yn), sd = sd(a$yn)) +
p[2] * dnorm(ys, mean = mean(b$yn), sd = sd(b$yn)))