我有一份数据,看起来像这样(实际数据有4000行):
tt <- structure(list(X = c("k__Fungi;p__Ascomycota;c__Pezizomycotina_cls_Incertae_sedis;o__Pezizomycotina_ord_Incertae_sedis;f__Pezizomycotina_fam_Incertae_sedis;g__Vermispora",
"k__Fungi;p__Ascomycota;c__Saccharomycetes;o__Saccharomycetales;f__Saccharomycetales_fam_Incertae_sedis;g__Candida",
"k__Fungi;p__Ascomycota;c__Sordariomycetes;o__Branch06;f__unidentified;g__unidentified",
"k__Bacteria;p__Proteobacteria;c__Betaproteobacteria;o__Burkholderiales;f__Oxalobacteraceae;g__Massilia",
"k__Bacteria;p__Actinobacteria;c__Actinobacteria;o__Actinomycetales;f__Pseudonocardiaceae;g__Amycolatopsis",
"k__Bacteria;p__Armatimonadetes;c__Armatimonadia;o__FW68;f__;g__",
"k__Bacteria;p__Proteobacteria;c__Deltaproteobacteria;o__Syntrophobacterales;f__Syntrophobacteraceae;g__",
"k__Fungi;p__Ascomycota;c__Dothideomycetes;o__Pleosporales;f__Leptosphaeriaceae;g__Leptosphaeria",
"D_0__Bacteria;D_1__WPS-2;D_2__metagenome;D_3__metagenome;D_4__metagenome;D_5__metagenome",
"D_0__Bacteria;D_1__Patescibacteria;D_2__Saccharimonadia;D_3__Saccharimonadales;D_4__Saccharimonadaceae;D_5__Candidatus Saccharibacteria bacterium RAAC3_TM7_1",
"D_0__Bacteria;D_1__Chloroflexi;D_2__AD3;D_3__uncultured bacterium;D_4__uncultured bacterium;D_5__uncultured bacterium",
"k__Fungi;p__Chytridiomycota;c__Chytridiomycetes;o__Rhizophydiales;f__Rhizophydiales_fam_Incertae_sedis;g__Coralloidiomyces",
"k__Bacteria;p__Bacteroidetes;c__Cytophagia;o__Cytophagales;f__Cytophagaceae;g__Pontibacter",
"D_0__Bacteria;D_1__WS2;D_2__uncultured soil bacterium;D_3__uncultured soil bacterium;D_4__uncultured soil bacterium;D_5__uncultured soil bacterium",
"k__Fungi;p__Basidiomycota;c__Agaricomycetes;o__Thelephorales;f__Thelephoraceae;g__Amaurodon",
"D_0__Bacteria;D_1__Patescibacteria;D_2__Parcubacteria;D_3__Candidatus Kaiserbacteria;D_4__metagenome;D_5__metagenome",
"k__Bacteria;p__Proteobacteria;c__Betaproteobacteria;o__Burkholderiales;f__Oxalobacteraceae;g__Janthinobacterium",
"k__Bacteria;p__Bacteroidetes;c__Bacteroidia;o__Bacteroidales;__;__",
"D_0__Bacteria;D_1__Bacteroidetes;D_2__Bacteroidia;D_3__Sphingobacteriales;D_4__Sphingobacteriaceae;D_5__Pedobacter",
"k__Bacteria;p__Acidobacteria;c__Acidobacteriia;o__Acidobacteriales;f__Koribacteraceae;g__",
"D_0__Bacteria;D_1__Planctomycetes;D_2__Planctomycetacia;D_3__Pirellulales;D_4__Pirellulaceae;D_5__Pirellula",
"k__Fungi;p__Ascomycota;c__Pezizomycetes;o__Pezizales;f__Sarcosomataceae;g__Urnula",
"D_0__Bacteria;D_1__Proteobacteria;D_2__Alphaproteobacteria;D_3__Azospirillales;D_4__uncultured;D_5__uncultured bacterium",
"k__Fungi;p__Basidiomycota;c__Agaricomycetes;o__Polyporales;f__Meruliaceae;g__Uncobasidium",
"D_0__Bacteria;D_1__Dependentiae;D_2__Babeliae;D_3__Babeliales;D_4__Vermiphilaceae;__",
"D_0__Bacteria;D_1__Proteobacteria;D_2__Alphaproteobacteria;D_3__Caulobacterales;D_4__Caulobacteraceae;D_5__Caulobacter",
"k__Fungi;p__Mortierellomycota;__;__;__;__", "k__Fungi;p__Ascomycota;c__Dothideomycetes;o__Pleosporales;f__Dictyosporiaceae;g__Dictyosporium",
"k__Bacteria;p__Acidobacteria;c__;o__;f__;g__", "D_0__Bacteria;D_1__Patescibacteria;D_2__Gracilibacteria;D_3__Absconditabacteriales (SR1);__;__",
"k__Fungi;p__Basidiomycota;c__Agaricomycetes;o__Agaricales;f__Entolomataceae;g__Clitopilus",
"k__Bacteria;p__Proteobacteria;c__Betaproteobacteria;o__Burkholderiales;f__Burkholderiaceae;g__",
"D_0__Bacteria;D_1__Armatimonadetes;D_2__Fimbriimonadia;D_3__Fimbriimonadales;D_4__Fimbriimonadaceae;D_5__Armatimonadetes bacterium 55-13",
"k__Fungi;p__Basidiomycota;c__Agaricomycetes;o__Agaricales;f__Bolbitiaceae;g__Agrocybe",
"k__Fungi;p__Ascomycota;c__Leotiomycetes;o__Helotiales;f__Helotiales_fam_Incertae_sedis;__",
"D_0__Bacteria;D_1__Cyanobacteria;D_2__Oxyphotobacteria;D_3__Oxyphotobacteria Incertae Sedis;D_4__Unknown Family;D_5__Leptolyngbya ANT.L52.2",
"k__Bacteria;p__Chloroflexi;c__Chloroflexi;o__[Roseiflexales];f__[Kouleothrixaceae];__",
"D_0__Bacteria;D_1__Proteobacteria;D_2__Alphaproteobacteria;D_3__Rhizobiales;D_4__Hyphomicrobiaceae;__",
"k__Fungi;p__Ascomycota;c__Eurotiomycetes;o__Onygenales;f__Onygenales_fam_Incertae_sedis;g__Spiromastix",
"k__Fungi;p__Basidiomycota;c__Tremellomycetes;o__Tremellales;f__Bulleribasidiaceae;g__Vishniacozyma",
"k__Fungi;p__Basidiomycota;c__Tremellomycetes;o__Tremellales;f__Bulleribasidiaceae;g__Bulleribasidium",
"k__Fungi;p__Ascomycota;c__Dothideomycetes;o__Pleosporales;f__Corynesporascaceae;g__Corynespora",
"k__Fungi;p__Ascomycota;c__Eurotiomycetes;o__Chaetothyriales;f__Cyphellophoraceae;g__Cyphellophora",
"k__Bacteria;p__Proteobacteria;c__Gammaproteobacteria;o__Xanthomonadales;f__Sinobacteraceae;__",
"D_0__Bacteria;D_1__Proteobacteria;D_2__Deltaproteobacteria;D_3__SAR324 clade(Marine group B);__;__",
"k__Fungi;p__Ascomycota;c__Leotiomycetes;o__Leotiomycetes_ord_Incertae_sedis;f__Myxotrichaceae;g__Oidiodendron",
"k__Bacteria;p__Proteobacteria;c__Deltaproteobacteria;o__Myxococcales;f__Cystobacteraceae;g__Cystobacter",
"D_0__Bacteria;D_1__Patescibacteria;D_2__Parcubacteria;D_3__Candidatus Kaiserbacteria;D_4__uncultured bacterium;D_5__uncultured bacterium",
"k__Fungi;p__Chytridiomycota;c__Spizellomycetes;o__Spizellomycetales;f__Spizellomycetaceae;g__Spizellomyces",
"k__Bacteria;p__Armatimonadetes;c__Chthonomonadetes;o__SJA-22;f__;g__"
), Reject.null.hypothesis = c("False", "False", "False", "False",
"False", "False", "False", "False", "False", "False", "False",
"False", "False", "False", "False", "False", "False", "False",
"False", "False", "False", "False", "False", "False", "False",
"False", "False", "False", "False", "False", "False", "False",
"False", "False", "False", "False", "False", "False", "False",
"False", "False", "False", "False", "False", "False", "False",
"False", "False", "False", "False"), W = c(3L, 57L, 89L, 0L,
3L, 0L, 6L, 33L, 27L, 28L, 1L, 3L, 5L, 1L, 3L, 1L, 0L, 10L, 1L,
15L, 26L, 1L, 1L, 4L, 1L, 0L, 6L, 15L, 0L, 23L, 6L, 1L, 45L,
4L, 0L, 1L, 5L, 1L, 45L, 3L, 7L, 192L, 53L, 0L, 2L, 8L, 0L, 18L,
7L, 0L), Type = structure(c(4L, 6L, 6L, 2L, 1L, 1L, 2L, 9L, 5L,
5L, 7L, 4L, 1L, 7L, 3L, 7L, 1L, 1L, 8L, 1L, 5L, 4L, 7L, 9L, 8L,
7L, 10L, 10L, 1L, 5L, 10L, 2L, 5L, 9L, 4L, 7L, 1L, 8L, 6L, 9L,
9L, 4L, 6L, 1L, 7L, 3L, 1L, 5L, 9L, 2L), .Label = c("Soil_16S_Monmouth_ancom.csv",
"Soil_16S_Urbana_ancom.csv", "Soil_ITS_Monmouth_ancom.csv", "Soil_ITS_urbana_ancom.csv",
"Rhizoshpere_16S_U_and_M_together_ancom.tsv", "Rhizoshpere_ITS_U_and_M_together_ancom.tsv",
"Rhizoshpere_16S_Monmouth_only_ancom.tsv", "Rhizoshpere_16S_Urbana_only_ancom.tsv",
"Rhizoshpere_ITS_Monmouth_only_ancom.tsv", "Rhizoshpere_ITS_Urbana_only_ancom.tsv"
), class = "factor"), microbiome_type = c("Soil", "Rhizosphere",
"Rhizosphere", "Soil", "Soil", "Soil", "Soil", "Rhizosphere",
"Rhizosphere", "Rhizosphere", "Rhizosphere", "Soil", "Soil",
"Rhizosphere", "Soil", "Rhizosphere", "Soil", "Soil", "Rhizosphere",
"Soil", "Rhizosphere", "Soil", "Rhizosphere", "Rhizosphere",
"Rhizosphere", "Rhizosphere", "Rhizosphere", "Rhizosphere", "Soil",
"Rhizosphere", "Rhizosphere", "Soil", "Rhizosphere", "Rhizosphere",
"Soil", "Rhizosphere", "Soil", "Rhizosphere", "Rhizosphere",
"Rhizosphere", "Rhizosphere", "Soil", "Rhizosphere", "Soil",
"Rhizosphere", "Soil", "Soil", "Rhizosphere", "Rhizosphere",
"Soil")), row.names = c(3771L, 3792L, 3806L, 2839L, 1913L, 2004L,
2956L, 3311L, 1619L, 1008L, 535L, 4785L, 2087L, 1625L, 4564L,
977L, 2836L, 2059L, 510L, 1706L, 1090L, 3750L, 1143L, 4514L,
848L, 1158L, 4873L, 3268L, 1658L, 952L, 4299L, 2792L, 408L, 4279L,
3610L, 827L, 2217L, 1275L, 3573L, 4713L, 4704L, 3258L, 3499L,
3018L, 1487L, 3668L, 2913L, 982L, 4806L, 2013L), class = "data.frame")
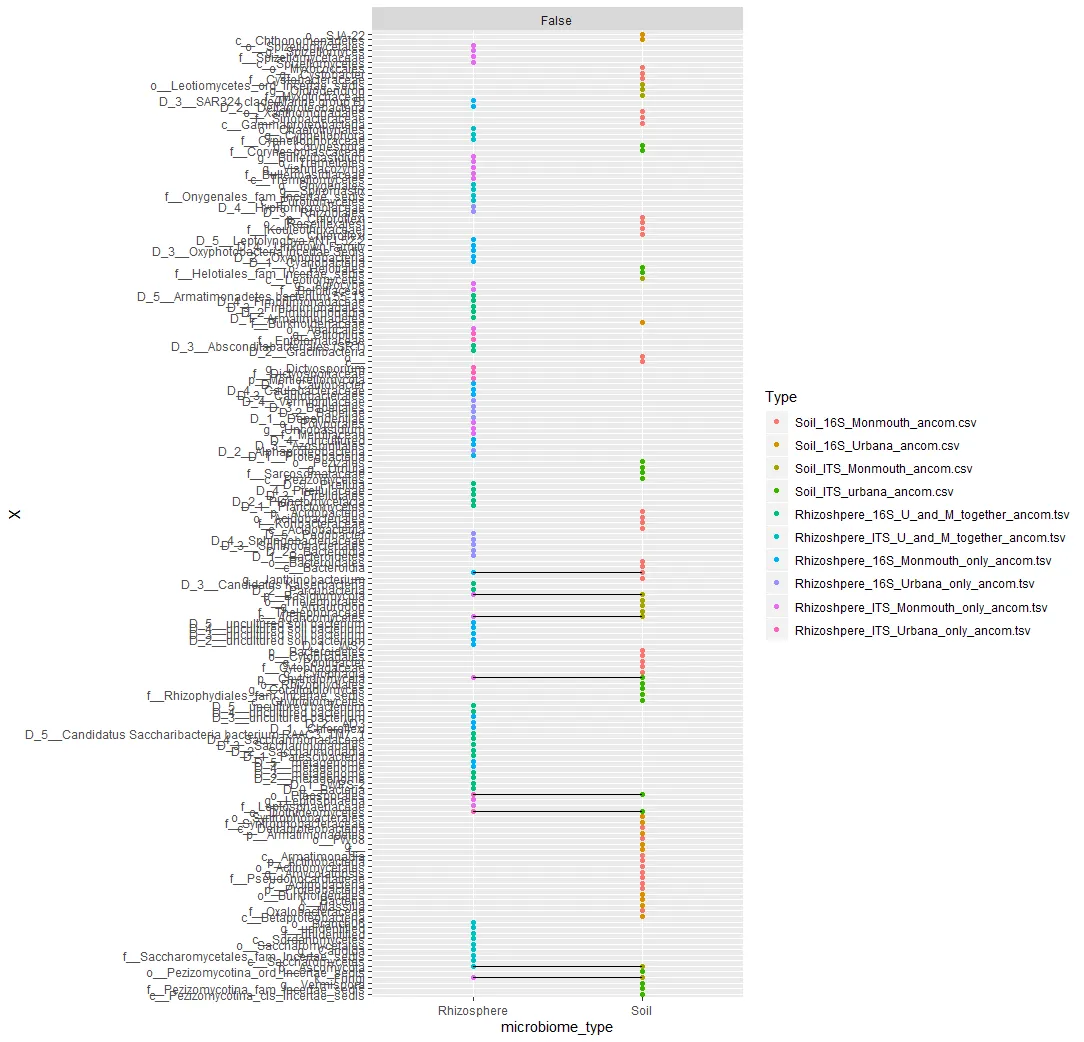
我想绘制
X与微生物群落类型的图表,以显示在土壤和根际中共同存在的X。也就是说,我想查看哪些细菌和真菌群落(来自列X)在Rhizosphere和Soil微生物群落类型之间被共享(以及未被共享)。我尝试了以下类似的内容,但它没有给我想要的结果:
ggplot(tt, aes(microbiome_type, X)) +
geom_jitter(aes(color = Type), size = 0.9)+
ggpubr::color_palette("jco")+
ggpubr::theme_pubclean() +
theme(axis.text.y=element_blank()) +
facet_wrap(~Reject.null.hypothesis)
请问有没有更好的方法来绘制这种类型的数据?