我在使用ggplot图例时遇到了一些问题,以下是我的第一段代码,只涉及到corrGenes的图例,这部分代码没有问题。
gene1=c(1.041,0.699,0.602,0.602,2.585,0.602,1.000,0.602,1.230,1.176,0.699,0.477,1.322)
BIME = c(0.477,0.477,0.301,0.477,2.398,0.301,0.602,0.301,0.602,0.699,0.602,0.477,1.176)
corrGenes=c(0.922,0.982,0.934,0.917,0.993,0.697,0.000,0.440,0.859,0.788,0.912,0.687,0.894)
DF=data.frame(gene1,BIME,corrGenes)
plot= ggplot(data=DF,aes(x=gene1,y=BIME))+
geom_point(aes(colour=corrGenes),size=5)+
ylab("BIME normalized counts (log10(RPKM))")+
xlab("gene1 normalized counts (log10(RPKM))")
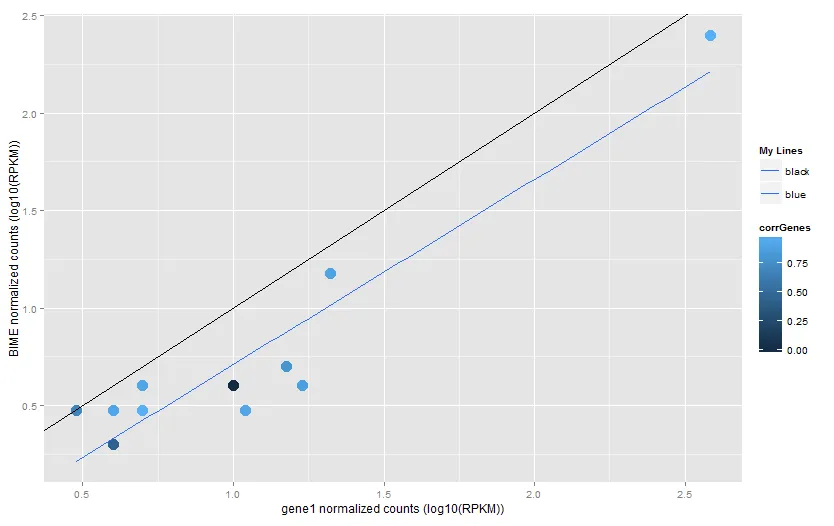
当我添加abline和smooth时,我会得到正确的图表:
plot= ggplot(data=DF,aes(x=gene1,y=BIME))+
geom_point(aes(colour=corrGenes),size=5)+
geom_abline(intercept=0, slope=1)+
stat_smooth(method = "lm",se=FALSE)+
ylab("BIME normalized counts (log10(RPKM))")+
xlab("gene1 normalized counts (log10(RPKM))")
但是没有办法获取它们的传说,我尝试了许多其他组合:
plot= ggplot(data=DF,aes(x=gene1,y=BIME))+
geom_point(aes(colour=corrGenes),size=5)+
geom_abline(aes(colour="best"),intercept=0, slope=1)+
stat_smooth(aes(colour="data"),method = "lm",se=FALSE)+
scale_colour_manual(name="Fit", values=c("data"="blue", "best"="black"))+
ylab("BIME normalized counts (log10(RPKM))")+
xlab("gene1 normalized counts (log10(RPKM))")
如果有人有解决这个微小但非常烦人的问题的想法,那将非常有帮助!
Error: Continuous value supplied to discrete scale。 - MesmerError: Continuous value supplied to discrete scale大多数情况下意味着R将你提供的字符型 x 值读取为连续值。尝试使用x <- as.factor(x)然后进行绘图。 - gented