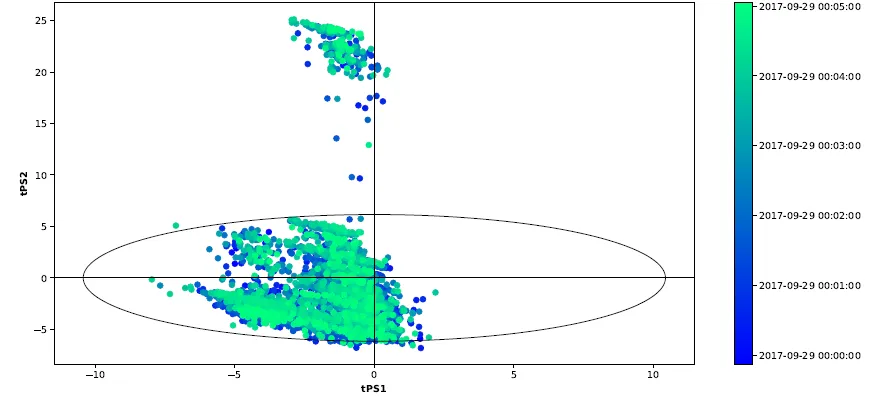
使用Hotelling's T2计算置信区间的Python PCA图
8
- Dr.Who
2
请查看以下链接:https://dev59.com/6mIj5IYBdhLWcg3wklyU - kvr
也可以尝试这个链接:https://dev59.com/S2ct5IYBdhLWcg3wKqUd - kvr
2个回答
6
这让我很困扰,所以我采用了PCA 和 Hotelling's T^2 for confidence intervall in R的答案,并在Python中使用了一些ggbiplot R包的源代码。
from sklearn import decomposition
from sklearn.preprocessing import StandardScaler
import numpy as np
import matplotlib.pyplot as plt
import scipy, random
#Generate data and fit PCA
random.seed(1)
data = np.array(np.random.normal(0, 1, 500)).reshape(100, 5)
outliers = np.array(np.random.uniform(5, 10, 25)).reshape(5, 5)
data = np.vstack((data, outliers))
pca = decomposition.PCA(n_components = 2)
scaler = StandardScaler()
scaler.fit(data)
data = scaler.transform(data)
pcaFit = pca.fit(data)
dataProject = pcaFit.transform(data)
#Calculate ellipse bounds and plot with scores
theta = np.concatenate((np.linspace(-np.pi, np.pi, 50), np.linspace(np.pi, -np.pi, 50)))
circle = np.array((np.cos(theta), np.sin(theta)))
sigma = np.cov(np.array((dataProject[:, 0], dataProject[:, 1])))
ed = np.sqrt(scipy.stats.chi2.ppf(0.95, 2))
ell = np.transpose(circle).dot(np.linalg.cholesky(sigma) * ed)
a, b = np.max(ell[: ,0]), np.max(ell[: ,1]) #95% ellipse bounds
t = np.linspace(0, 2 * np.pi, 100)
plt.scatter(dataProject[:, 0], dataProject[:, 1])
plt.plot(a * np.cos(t), b * np.sin(t), color = 'red')
plt.grid(color = 'lightgray', linestyle = '--')
plt.show()
- srhoades10
5
pca库提供Hotelling T2和SPE/DmodX异常点检测功能。
pip install pca
from pca import pca
import pandas as pd
import numpy as np
# Create dataset with 100 samples
X = np.array(np.random.normal(0, 1, 500)).reshape(100, 5)
# Create 5 outliers
outliers = np.array(np.random.uniform(5, 10, 25)).reshape(5, 5)
# Combine data
X = np.vstack((X, outliers))
# Initialize model. Alpha is the threshold for the hotellings T2 test to determine outliers in the data.
model = pca(alpha=0.05)
# Fit transform
out = model.fit_transform(X)
使用以下代码打印异常值
print(out['outliers'])
# y_proba y_score y_bool y_bool_spe y_score_spe
# 1.0 9.799576e-01 3.060765 False False 0.993407
# 1.0 8.198524e-01 5.945125 False False 2.331705
# 1.0 9.793117e-01 3.086609 False False 0.128518
# 1.0 9.743937e-01 3.268052 False False 0.794845
# 1.0 8.333778e-01 5.780220 False False 1.523642
# .. ... ... ... ... ...
# 1.0 6.793085e-11 69.039523 True True 14.672828
# 1.0 2.610920e-291 1384.158189 True True 16.566568
# 1.0 6.866703e-11 69.015237 True True 14.936442
# 1.0 1.765139e-292 1389.577522 True True 17.183093
# 1.0 1.351102e-291 1385.483398 True True 17.319038
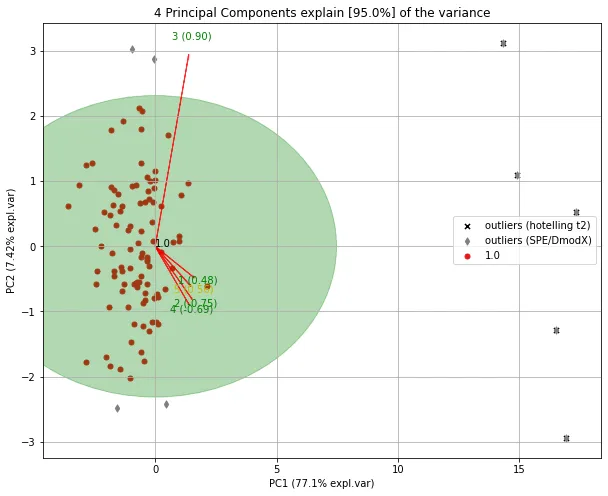
制作图表。
model.biplot(legend=True, SPE=True, hotellingt2=True)
- erdogant
网页内容由stack overflow 提供, 点击上面的可以查看英文原文,
原文链接
原文链接