我有一些嘈杂的数据,正在尝试制作高低包络线到信号。这有点像MATLAB中“提取峰值包络”的此示例。
Python中是否有类似的函数能够实现呢?我的整个项目都是用Python编写的,最坏的情况是我可以提取numpy数组并将其放入MATLAB中使用该示例。但我更喜欢matplotlib的外观... 真的很烦做所有的MATLAB和Python之间的I/O...
我有一些嘈杂的数据,正在尝试制作高低包络线到信号。这有点像MATLAB中“提取峰值包络”的此示例。
Python中是否有类似的函数能够实现呢?我的整个项目都是用Python编写的,最坏的情况是我可以提取numpy数组并将其放入MATLAB中使用该示例。但我更喜欢matplotlib的外观... 真的很烦做所有的MATLAB和Python之间的I/O...
import numpy as np
from matplotlib import pyplot as plt
def hl_envelopes_idx(s, dmin=1, dmax=1, split=False):
"""
Input :
s: 1d-array, data signal from which to extract high and low envelopes
dmin, dmax: int, optional, size of chunks, use this if the size of the input signal is too big
split: bool, optional, if True, split the signal in half along its mean, might help to generate the envelope in some cases
Output :
lmin,lmax : high/low envelope idx of input signal s
"""
# locals min
lmin = (np.diff(np.sign(np.diff(s))) > 0).nonzero()[0] + 1
# locals max
lmax = (np.diff(np.sign(np.diff(s))) < 0).nonzero()[0] + 1
if split:
# s_mid is zero if s centered around x-axis or more generally mean of signal
s_mid = np.mean(s)
# pre-sorting of locals min based on relative position with respect to s_mid
lmin = lmin[s[lmin]<s_mid]
# pre-sorting of local max based on relative position with respect to s_mid
lmax = lmax[s[lmax]>s_mid]
# global min of dmin-chunks of locals min
lmin = lmin[[i+np.argmin(s[lmin[i:i+dmin]]) for i in range(0,len(lmin),dmin)]]
# global max of dmax-chunks of locals max
lmax = lmax[[i+np.argmax(s[lmax[i:i+dmax]]) for i in range(0,len(lmax),dmax)]]
return lmin,lmax
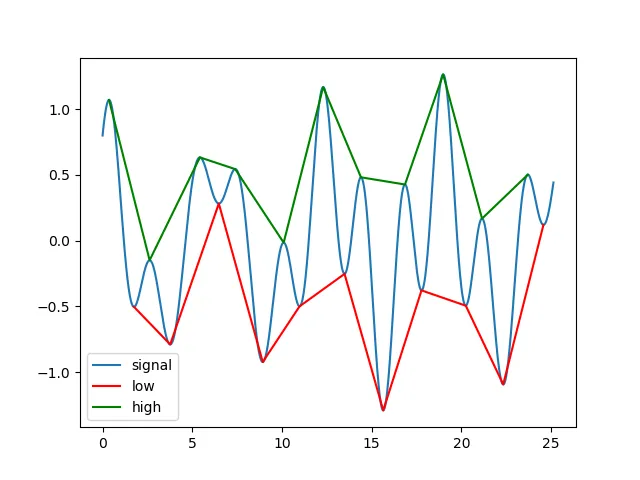
例子1:准周期振动
t = np.linspace(0,8*np.pi,5000)
s = 0.8*np.cos(t)**3 + 0.5*np.sin(np.exp(1)*t)
lmin, lmax = hl_envelopes_idx(s)
# plot
plt.plot(t,s,label='signal')
plt.plot(t[lmin], s[lmin], 'r', label='low')
plt.plot(t[lmax], s[lmax], 'g', label='high')
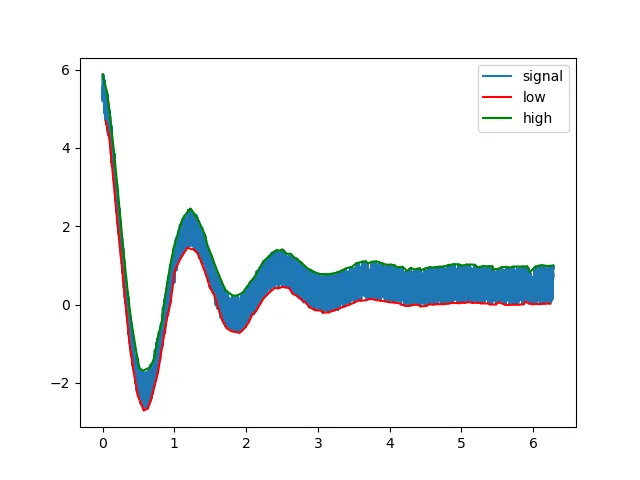
示例2:嘈杂衰减信号
t = np.linspace(0,2*np.pi,5000)
s = 5*np.cos(5*t)*np.exp(-t) + np.random.rand(len(t))
lmin, lmax = hl_envelopes_idx(s,dmin=15,dmax=15)
# plot
plt.plot(t,s,label='signal')
plt.plot(t[lmin], s[lmin], 'r', label='low')
plt.plot(t[lmax], s[lmax], 'g', label='high')
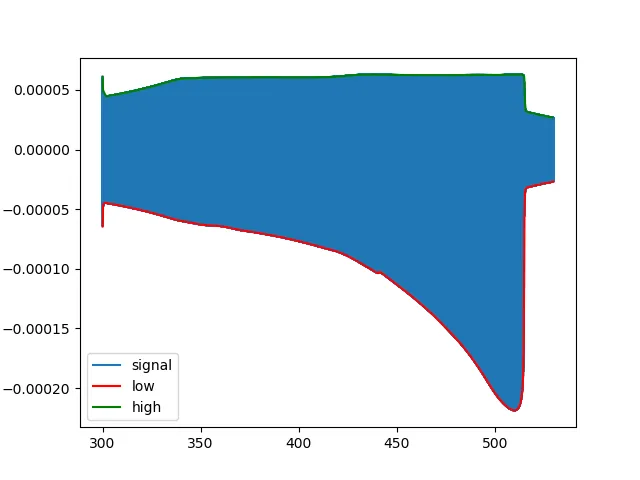
示例3:非对称调制啁啾信号
一个更加复杂的信号,包含18867925个样本(此处未包含):
from numpy import array, sign, zeros
from scipy.interpolate import interp1d
from matplotlib.pyplot import plot,show,hold,grid
s = array([1,4,3,5,3,2,4,3,4,5,4,3,2,5,6,7,8,7,8]) #This is your noisy vector of values.
q_u = zeros(s.shape)
q_l = zeros(s.shape)
#Prepend the first value of (s) to the interpolating values. This forces the model to use the same starting point for both the upper and lower envelope models.
u_x = [0,]
u_y = [s[0],]
l_x = [0,]
l_y = [s[0],]
#Detect peaks and troughs and mark their location in u_x,u_y,l_x,l_y respectively.
for k in xrange(1,len(s)-1):
if (sign(s[k]-s[k-1])==1) and (sign(s[k]-s[k+1])==1):
u_x.append(k)
u_y.append(s[k])
if (sign(s[k]-s[k-1])==-1) and ((sign(s[k]-s[k+1]))==-1):
l_x.append(k)
l_y.append(s[k])
#Append the last value of (s) to the interpolating values. This forces the model to use the same ending point for both the upper and lower envelope models.
u_x.append(len(s)-1)
u_y.append(s[-1])
l_x.append(len(s)-1)
l_y.append(s[-1])
#Fit suitable models to the data. Here I am using cubic splines, similarly to the MATLAB example given in the question.
u_p = interp1d(u_x,u_y, kind = 'cubic',bounds_error = False, fill_value=0.0)
l_p = interp1d(l_x,l_y,kind = 'cubic',bounds_error = False, fill_value=0.0)
#Evaluate each model over the domain of (s)
for k in xrange(0,len(s)):
q_u[k] = u_p(k)
q_l[k] = l_p(k)
#Plot everything
plot(s);hold(True);plot(q_u,'r');plot(q_l,'g');grid(True);show()
上述代码未过滤可能发生在某个阈值“距离”(如时间)内的峰值或谷值。这类似于envelope的第二个参数。然而,通过检查u_x,u_y连续值之间的差异来轻松添加它。
然而,先使用移动平均滤波器对数据进行低通滤波可以快速改善前面提到的问题,然后再插值上下包络函数。您可以通过将(s)与适当的移动平均滤波器卷积来轻松实现此操作。不在此处详细介绍(如果需要可以提供),但为了生成一个该滤波器可以对N个连续样本起作用的移动平均滤波器,您可以执行以下操作:s_filtered = numpy.convolve(s, numpy.ones((1,N))/float(N) 。 (N)越高,您的数据看起来就越平稳。请注意,由于称为group delay的平滑滤波器的原因,在s_filtered中,这将使(s)值向右移动(N/2)个样本。有关移动平均值的更多信息,请参见this link 。
在@A_A的回答基础上,用nim/max测试来代替符号检查,使代码更加健壮。
import numpy as np
import scipy.interpolate
import matplotlib.pyplot as pt
%matplotlib inline
t = np.multiply(list(range(1000)), .1)
s = 10*np.sin(t)*t**.5
u_x = [0]
u_y = [s[0]]
l_x = [0]
l_y = [s[0]]
#Detect peaks and troughs and mark their location in u_x,u_y,l_x,l_y respectively.
for k in range(2,len(s)-1):
if s[k] >= max(s[:k-1]):
u_x.append(t[k])
u_y.append(s[k])
for k in range(2,len(s)-1):
if s[k] <= min(s[:k-1]):
l_x.append(t[k])
l_y.append(s[k])
u_p = scipy.interpolate.interp1d(u_x, u_y, kind = 'cubic', bounds_error = False, fill_value=0.0)
l_p = scipy.interpolate.interp1d(l_x, l_y, kind = 'cubic', bounds_error = False, fill_value=0.0)
q_u = np.zeros(s.shape)
q_l = np.zeros(s.shape)
for k in range(0,len(s)):
q_u[k] = u_p(t[k])
q_l[k] = l_p(t[k])
pt.plot(t,s)
pt.plot(t, q_u, 'r')
pt.plot(t, q_l, 'g')
如果你期望函数是递增的,请尝试:
最初的回答:
for k in range(1,len(s)-2):
if s[k] <= min(s[k+1:]):
l_x.append(t[k])
l_y.append(s[k])
用于下凸壳。
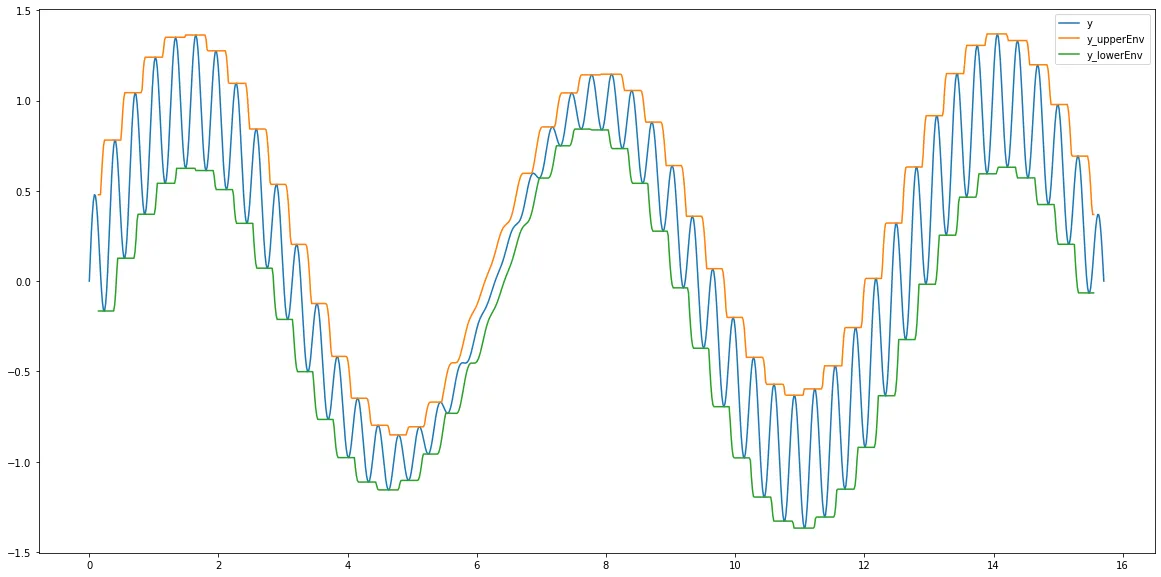
或者您可以使用pandas。这里我只需要两行代码:
import pandas as pd
import numpy as np
x=np.linspace(0,5*np.pi,1000)
y=np.sin(x)+0.4*np.cos(x/4)*np.sin(x*20)
df=pd.DataFrame(data={"y":y},index=x)
windowsize = 20
df["y_upperEnv"]=df["y"].rolling(window=windowsize).max().shift(int(-windowsize/2))
df["y_lowerEnv"]=df["y"].rolling(window=windowsize).min().shift(int(-windowsize/2))
df.plot(figsize=(20,10))
Output:
def envelope(sig, distance):
# split signal into negative and positive parts
u_x = np.where(sig > 0)[0]
l_x = np.where(sig < 0)[0]
u_y = sig.copy()
u_y[l_x] = 0
l_y = -sig.copy()
l_y[u_x] = 0
# find upper and lower peaks
u_peaks, _ = scipy.signal.find_peaks(u_y, distance=distance)
l_peaks, _ = scipy.signal.find_peaks(l_y, distance=distance)
# use peaks and peak values to make envelope
u_x = u_peaks
u_y = sig[u_peaks]
l_x = l_peaks
l_y = sig[l_peaks]
# add start and end of signal to allow proper indexing
end = len(sig)
u_x = np.concatenate((u_x, [0, end]))
u_y = np.concatenate((u_y, [0, 0]))
l_x = np.concatenate((l_x, [0, end]))
l_y = np.concatenate((l_y, [0, 0]))
# create envelope functions
u = scipy.interpolate.interp1d(u_x, u_y)
l = scipy.interpolate.interp1d(l_x, l_y)
return u, l
def test():
x = np.arange(200)
sig = np.sin(x)
u, l = envelope(sig, 1)
plt.figure(figsize=(25,5))
plt.plot(x, u(x))
plt.plot(x, l(x))
plt.plot(x, sig*0.9)
plt.show()
test()