在CalibratedClassifierCV的文档中指出:“用于训练校准器的样本不应用于训练目标分类器。”据我理解,如果我们使用X_train来训练我们的模型,那么我们就不应该使用X_train来训练校准器(因为我认为它只是将X_train映射到model.predict_proba?)。但是,使用已用于超参数优化的验证集X_val来校准校准器是否公平呢?
使用相同的验证集对模型和CalibratedClassifierCV进行校准
3
- CutePoison
1
这个问题更适合在stats.SE或datascience.SE上提问。 - Ben Reiniger
1个回答
2
CalibratedClassifierCV会在一部分数据上拟合分类器,然后查看预测概率是否与另一个数据集上的真实标签相对应。然后它尝试调整(“校准”)概率,使得对于具有0.7概率的一组样本,将有大约0.7个真实标签为“1”(用于二元分类)。注意,训练分类器和拟合校准器是在不同的数据集上完成的,以考虑可能存在的偏差。这可以通过两种不同的方式完成:
1.
CalibratedClassifierCV(..., cv=None, ensemble=True)。根据提供的cv策略划分您的训练数据,例如cv=5。像往常一样,您在k-1个折叠上训练基础估计器(要校准的分类器),然后在第k个剩下的折叠上拟合校准器。测试的结果预测将是k个已拟合校准器的平均值(“集成”)。2. 如果
ensemble=False,则使用单次拟合的cross_val_predict来拟合校准器。有了这个理解:
使用已经用于超参数优化的验证集X_val来校准校准器是否公平?
您可以找到最佳的超参数,然后使用相同的训练数据集,但选择了CV策略(Ensemble True或False)来拟合校准器。请再次注意,您永远不会在同一数据上拟合分类器和校准器。
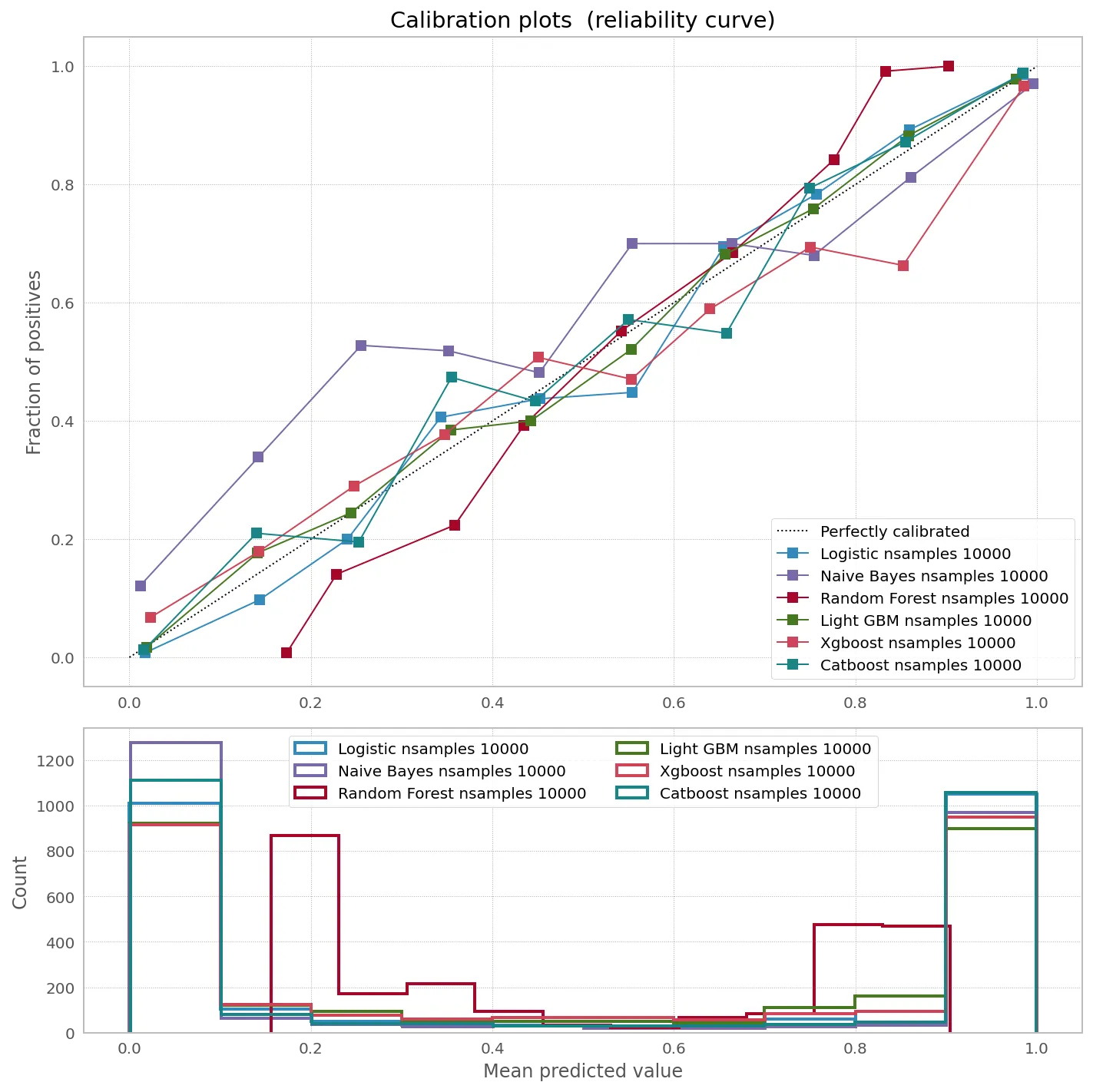
有趣的是,根据分类器和数据大小,校准可能会或可能不会帮助提高概率预测的准确性:
from sklearn import datasets
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.calibration import calibration_curve, CalibratedClassifierCV
from xgboost import XGBClassifier
from lightgbm import LGBMClassifier
from catboost import CatBoostClassifier
np.random.seed(42)
# Create classifiers
lrc = LogisticRegression(n_jobs=-1)
gnb = GaussianNB()
svc = SVC(C=1.0, probability=True,)
rfc = RandomForestClassifier(n_estimators=300, max_depth=3,n_jobs=-1)
xgb = XGBClassifier(
n_estimators=300,
max_depth=3,
objective="binary:logistic",
eval_metric="logloss",
use_label_encoder=False,
)
lgb = LGBMClassifier(n_estimators=300, objective="binary", max_depth=3)
cat = CatBoostClassifier(n_estimators=300, max_depth=3, objective="Logloss", verbose=0)
df = pd.DataFrame()
plt.figure(figsize=(10, 10))
ax1 = plt.subplot2grid((3, 1), (0, 0), rowspan=2)
ax2 = plt.subplot2grid((3, 1), (2, 0))
ax1.plot([0, 1], [0, 1], "k:", label="Perfectly calibrated")
for clf, name in [
(lrc, "Logistic"),
(gnb, "Naive Bayes"),
# (svc, "Support Vector Classification"),
(rfc, "Random Forest"),
(lgb, "Light GBM"),
(xgb, "Xgboost"),
(cat, "Catboost"),
]:
print(name)
for nsamples in [1000,10000,100000]:
train_samples = 0.75
X, y = make_classification(
n_samples=nsamples, n_features=20, n_informative=2, n_redundant=2
)
i = int(train_samples * nsamples)
X_train = X[:i]
X_test = X[i:]
y_train = y[:i]
y_test = y[i:]
clf.fit(X_train, y_train)
prob_pos = clf.predict_proba(X_test)[:, 1]
fraction_of_positives, mean_predicted_value = calibration_curve(
y_test, prob_pos, n_bins=10
)
if nsamples in [10000]:
ax1.plot(
mean_predicted_value,
fraction_of_positives,
"s-",
label="%s" % (name + " nsamples " + str(nsamples),),
)
ax2.hist(
prob_pos,
bins=10,
label="%s" % (name + " nsamples " + str(nsamples),),
histtype="step",
lw=2,
)
preds = clf.predict_proba(X_test)
ll_before = log_loss(y_test, preds)
preds = (
CalibratedClassifierCV(clf, cv=5)
.fit(X_train, y_train)
.predict_proba(X_test)
)
ll_after = log_loss(y_test, preds)
df = df.append(pd.DataFrame({
"Samples": [nsamples],
"Model": name,
"LogLoss Before": round(ll_before,4),
"LogLoss After": round(ll_after,4),
"Gain": round(ll_before/ll_after,4)
}))
ax1.set_ylabel("Fraction of positives")
ax1.set_ylim([-0.05, 1.05])
ax1.legend(loc="lower right")
ax1.set_title("Calibration plots (reliability curve)")
ax2.set_xlabel("Mean predicted value")
ax2.set_ylabel("Count")
ax2.legend(loc="upper center", ncol=2)
plt.tight_layout()
print(df)
Samples Model LogLoss Before LogLoss After Gain
0 1000 Logistic 0.3941 0.3854 1.0226
0 10000 Logistic 0.1340 0.1345 0.9959
0 100000 Logistic 0.1645 0.1645 0.9999
0 1000 Naive Bayes 0.3025 0.2291 1.3206
0 10000 Naive Bayes 0.4094 0.3055 1.3403
0 100000 Naive Bayes 0.4119 0.2594 1.5881
0 1000 Random Forest 0.4438 0.3137 1.4146
0 10000 Random Forest 0.3450 0.2776 1.2427
0 100000 Random Forest 0.3104 0.1642 1.8902
0 1000 Light GBM 0.2993 0.2219 1.3490
0 10000 Light GBM 0.2074 0.2182 0.9507
0 100000 Light GBM 0.2397 0.2534 0.9459
0 1000 Xgboost 0.1870 0.1638 1.1414
0 10000 Xgboost 0.3072 0.2967 1.0351
0 100000 Xgboost 0.1136 0.1186 0.9575
0 1000 Catboost 0.1834 0.1901 0.9649
0 10000 Catboost 0.1251 0.1377 0.9085
0 100000 Catboost 0.1600 0.1727 0.9264
- Sergey Bushmanov
网页内容由stack overflow 提供, 点击上面的可以查看英文原文,
原文链接
原文链接