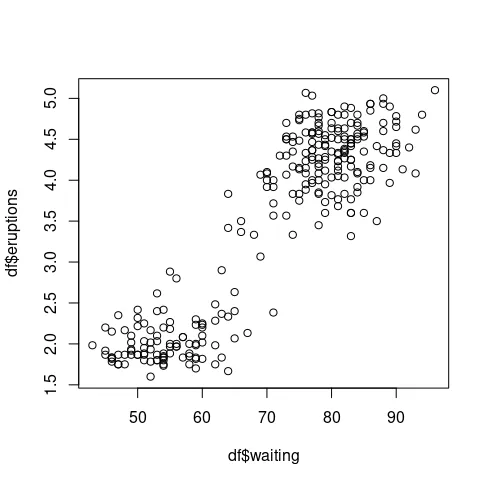
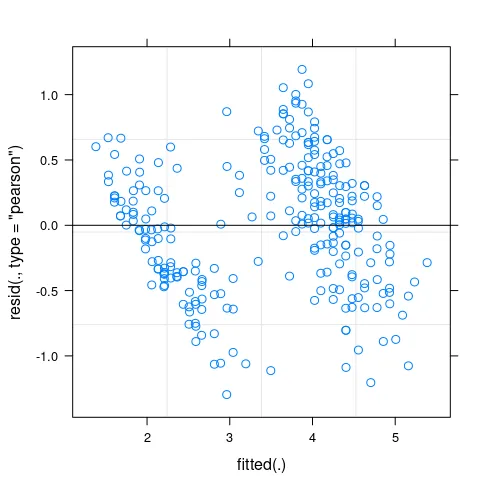
标题已经说明了一切: 在lme4中更改随机效应分组变量(例如重复测量实验中被试的名称)的(据说是任意的)标签可能会改变结果输出。最小示例:
但这个不行:
require(dplyr)
require(lme4)
require(digest)
df = faithful %>% mutate(subject = rep(as.character(1:8), each = 34),
subject2 = rep(as.character(9:16), each = 34))
summary(lmer(eruptions ~ waiting + (waiting | subject), data = df))$coefficients[2,1] # = 0.07564181
summary(lmer(eruptions ~ waiting + (waiting | subject2), data = df))$coefficients[2,1] # = 0.07567655
我认为这是由于lme4将它们转换为因子(factor),而不同的名称会产生不同的因子水平排序。例如,以下代码会导致问题:
df2 = faithful %>% mutate(subject = factor(rep(as.character(1:8), each = 34)),
subject2 = factor(rep(as.character(9:16), each = 34)))
summary(lmer(eruptions ~ waiting + (waiting | subject), data = df2))$coefficients[2,1] # = 0.07564181
summary(lmer(eruptions ~ waiting + (waiting | subject2), data = df2))$coefficients[2,1] # = 0.07567655
但这个不行:
df3 = faithful %>% mutate(subject = factor(rep(as.character(1:8), each = 34)),
subject2 = factor(rep(as.character(1:8), each = 34),
levels = as.character(1:8),
labels = as.character(9:16)))
summary(lmer(eruptions ~ waiting + (waiting | subject), data = df3))$coefficients[2,1] # = 0.07564181
summary(lmer(eruptions ~ waiting + (waiting | subject2), data = df3))$coefficients[2,1] # = 0.07564181
这似乎是lme4中的一个问题。不同的任意变量标签不应该产生不同的输出,对吧?我有遗漏什么吗?为什么lme4会这样做?
(我知道输出差异很小,但在其他情况下我得到了更大的差异,足以改变p值从0.055到0.045等。此外,如果这是正确的,我认为它可能会导致轻微的再现性问题--例如,如果实验者在完成分析后匿名化他们的人类主体数据(通过更改名称),然后将其发布在公共存储库中。)