我经常处理需要一些重编码的字符数据。一个常见场景是,一个被记录的字符向量本质上是一个因子,但并不一定是一个类别。例如,考虑以下 chr 向量 vec:
set.seed(2021)
vec <- sample(rep(c("animal_dog_xyz", "animal_cat_abc", "animal_alligator_tyl"), 10))
vec
#> [1] "animal_dog_xyz" "animal_alligator_tyl" "animal_cat_abc"
#> [4] "animal_cat_abc" "animal_alligator_tyl" "animal_alligator_tyl"
#> [7] "animal_cat_abc" "animal_cat_abc" "animal_cat_abc"
#> [10] "animal_dog_xyz" "animal_dog_xyz" "animal_cat_abc"
#> [13] "animal_alligator_tyl" "animal_alligator_tyl" "animal_alligator_tyl"
#> [16] "animal_cat_abc" "animal_dog_xyz" "animal_alligator_tyl"
#> [19] "animal_alligator_tyl" "animal_cat_abc" "animal_dog_xyz"
#> [22] "animal_cat_abc" "animal_cat_abc" "animal_dog_xyz"
#> [25] "animal_dog_xyz" "animal_dog_xyz" "animal_dog_xyz"
#> [28] "animal_dog_xyz" "animal_alligator_tyl" "animal_alligator_tyl"
本文创建于2021-07-19,使用reprex包(v2.0.0)
如果我想重新编码这个向量并提取出动物名称,我会选择适用于字符数据的解决方案:
library(stringr)
sapply(str_split(vec, "_", n = 3), `[`, 2)
#> [1] "dog" "alligator" "cat" "cat" "alligator" "alligator"
#> [7] "cat" "cat" "cat" "dog" "dog" "cat"
#> [13] "alligator" "alligator" "alligator" "cat" "dog" "alligator"
#> [19] "alligator" "cat" "dog" "cat" "cat" "dog"
#> [25] "dog" "dog" "dog" "dog" "alligator" "alligator"
问题
如果向量非常长,进行这样的重新编码过程需要很长时间。R会迭代每个向量元素并应用该过程。鉴于向量中只有3个唯一值,这似乎效率低下。换句话说,我们不需要逐个检查元素并确定重新编码的值。
在这里,vec_long 长度为30000。这是在我的计算机上重新编码所需的时间。
vec_long <- sample(rep(c("animal_dog_xyz", "animal_cat_abc", "animal_alligator_tyl"), 10000))
length(vec_long)
#> [1] 30000
library(microbenchmark)
microbenchmark(sapply(str_split(vec_long, "_", n = 3), `[`, 2))
#> Unit: milliseconds
#> expr min lq mean
#> sapply(str_split(vec_long, "_", n = 3), `[`, 2) 51.6972 52.66918 57.42299
#> median uq max neval
#> 54.47867 58.7653 115.754 100
有没有一种方法可以利用这个向量实际上是一个因子的事实?因此识别唯一值(“级别”),重新编码它们,并重新部署到整个向量长度中?是否有这样的过程可以加快处理时间?
谢谢!
编辑
我想总结一下基于@GKi的答案,@ThomasIsCoding的答案和@user20650的评论所做的测试。
## The Data
set.seed(2021)
unique_vals <- c("animal_dog_xyz", "animal_cat_abc", "animal_alligator_tyl")
vec <- sample(rep(unique_vals, 10))
vec_long <- sample(rep(unique_vals, 1000))
vec_very_long <- sample(unique_vals, 100000))
## The functions
## function #1 -- as @user20650 proposed
via_fac_levels <- function(x) {
x_factor <- factor(x)
levels(x_factor) <- sapply(str_split(levels(x_factor), "_", n = 3), `[`, 2)
as.character(x_factor)
}
####################
## function #2 -- as @GKi proposed
via_fac_no_levels <- function(x) {
x_factor <- as.factor(x)
x_factor <- sapply(strsplit(levels(x_factor), "_", TRUE), `[`, 2)[x_factor]
as.character(x_factor)
}
####################
## function #3 -- the original slow method shown in the question
via_chr_only <- function(x) {
sapply(str_split(x, "_", n = 3), `[`, 2)
}
####################
## function #4 -- as @ThomasIsCoding proposed
via_read_table <- function(x) {
read.table(text = paste0(x, collapse = "\n"), sep = "_", header = FALSE)$V2
}
###################
## function #5 -- forcats::fct_relabel()
via_fct_relabel <- function(x) {
x_factor <- as.factor(x)
x_factor <- fct_relabel(x_factor, ~sapply(str_split(.x, "_", n = 3), `[`, 2))
as.character(x_factor)
}
## Performance assessment
### I ran it on Rstudio cloud
bm_short <- bench::mark(fac_levels = via_fac_levels(vec),
fac_no_levels = via_fac_no_levels(vec),
chr = via_chr_only(vec),
read_t = via_read_table(vec),
fct_relabel = via_fct_relabel(vec),
iterations = 1000)
bm_long <- bench::mark(fac_levels = via_fac_levels(vec_long),
fac_no_levels = via_fac_no_levels(vec_long),
chr = via_chr_only(vec_long),
read_t = via_read_table(vec_long),
fct_relabel = via_fct_relabel(vec_long),
iterations = 1000)
bm_very_long <- bench::mark(fac_levels = via_fac_levels(vec_very_long),
fac_no_levels = via_fac_no_levels(vec_very_long),
chr = via_chr_only(vec_very_long),
read_t = via_read_table(vec_very_long),
fct_relabel = via_fct_relabel(vec_very_long),
iterations = 1000)
## visualize
library(ggplot2)
library(tidyr)
library(ggbeeswarm)
library(beeswarm)
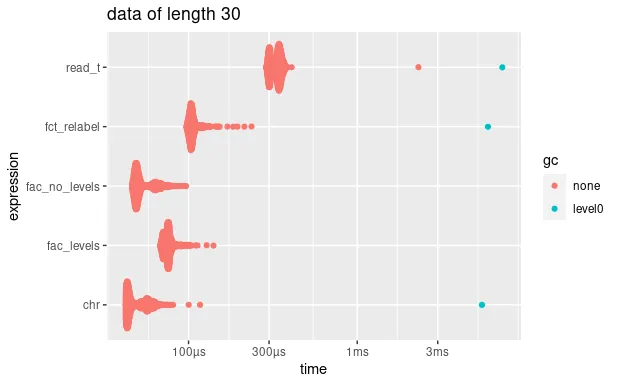
autoplot(bm_short) + ggtitle("data of length 30")
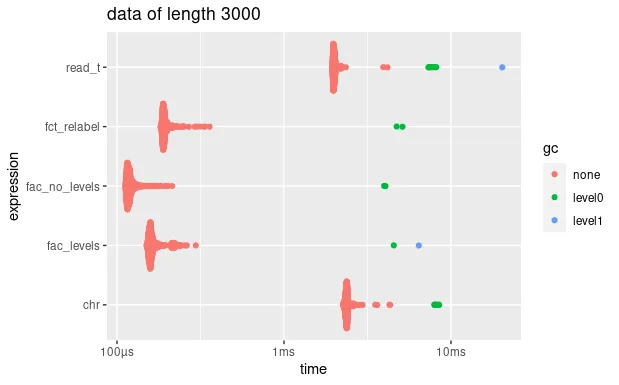
autoplot(bm_long) + ggtitle("data of length 3000")
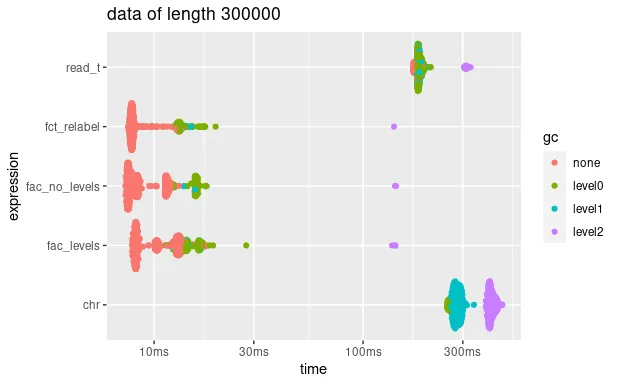
autoplot(bm_very_long) + ggtitle("data of length 300000")
## verify all functions give the same output
v1 <- via_fac_levels(vec_long)
v2 <- via_fac_no_levels(vec_long)
v3 <- via_chr_only(vec_long)
v4 <- via_read_table(vec_long)
v5 <- via_fct_relabel(vec_long)
all(sapply(list(v1, v2, v3, v4), FUN = identical, v5)) # https://dev59.com/p10Z5IYBdhLWcg3w-UV-#30850654
## [1] TRUE
vec_long = factor(vec_long) ; levels(vec_long) <- c("all", "cat", "dog")- user20650