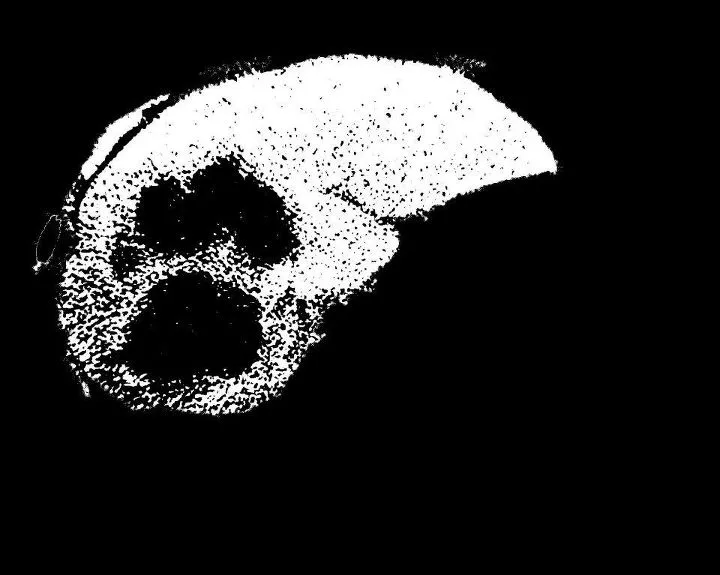
我有一个分段的肝脏。我需要分割其中的肿瘤。我使用了FCM方法。这是一个3级的FCM阈值分割。当我将其应用于图像时,我需要肿瘤区域(比其他部分更暗的区域)单独进行分割。但是我得到了相反的结果。周围所有的区域都被分割了。请帮忙解决。程序包括两个文件,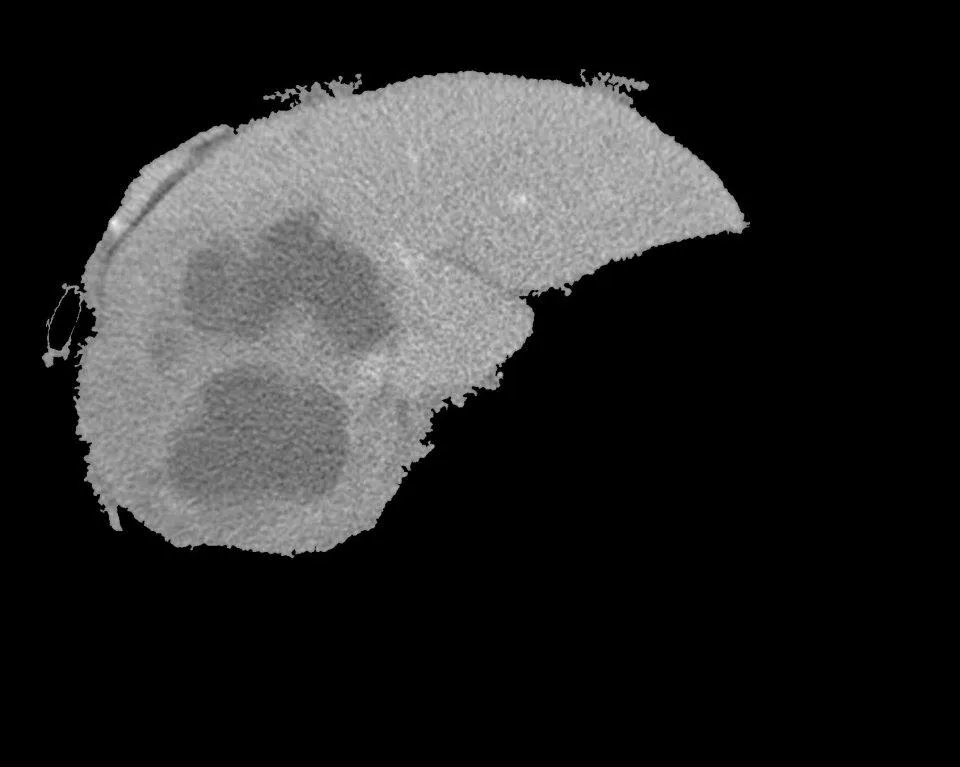
 从这里开始
我尝试使用
从这里开始
我尝试使用
testfcmthresh.m和一个函数fcmthresh.m
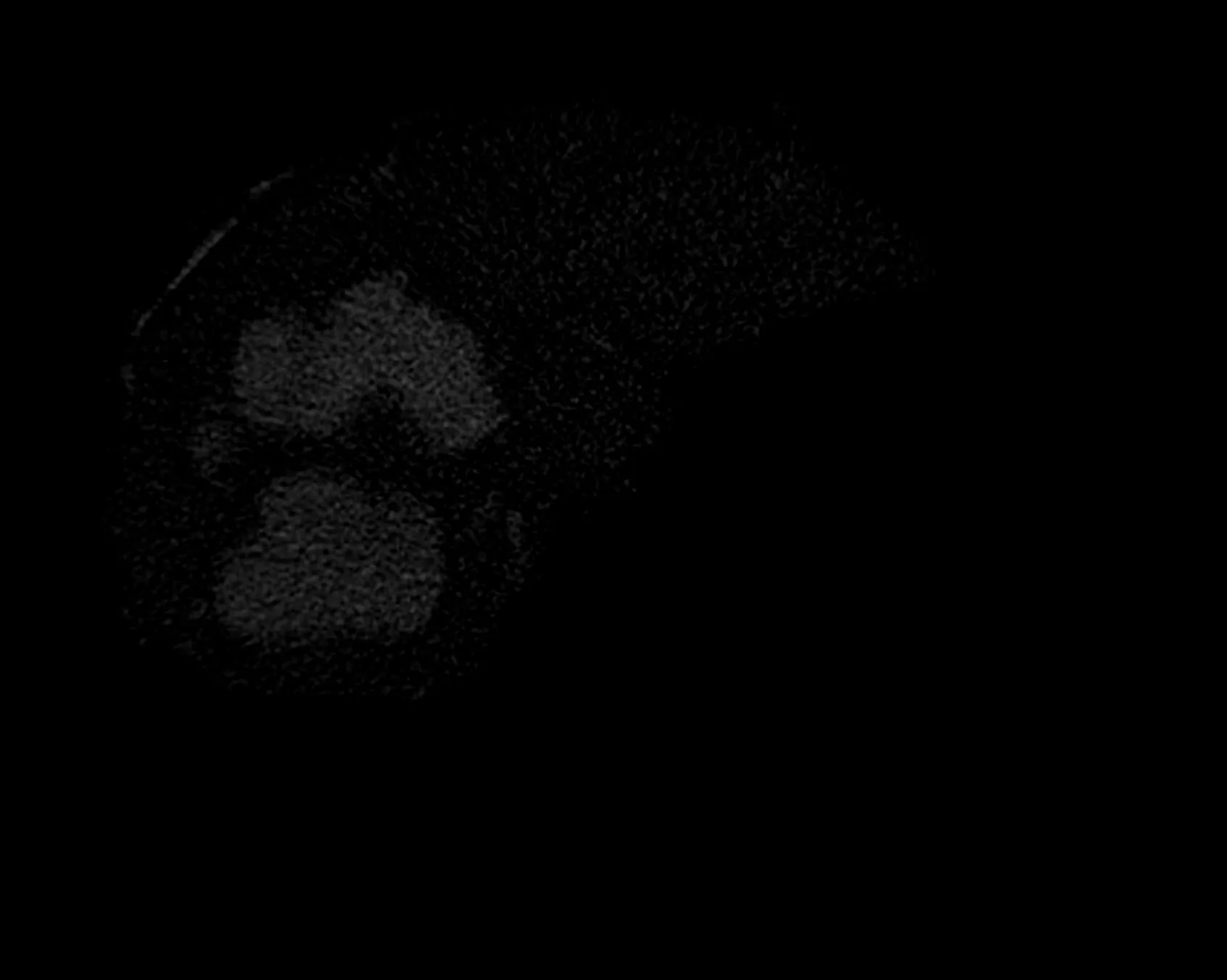
输入为“分段肝脏(使用区域生长)”和FCM输出图像:

 从这里开始
我尝试使用
从这里开始
我尝试使用imcomplement()补充图像。但是我得到的整个背景也变成了白色,因为背景原本很暗。请帮忙解决。

function [bw,level]=fcmthresh(IM,sw)
%FCMTHRESH Thresholding by 3-class fuzzy c-means clustering
% [bw,level]=fcmthresh(IM,sw) outputs the binary image bw and threshold level of
% image IM using a 3-class fuzzy c-means clustering. It often works better
% than Otsu's methold which outputs larger or smaller threshold on
% fluorescence images.
% sw is 0 or 1, a switch of cut-off position.
% sw=0, cut between the small and middle class
% sw=1, cut between the middle and large class
%
% Contributed by Guanglei Xiong (xgl99@mails.tsinghua.edu.cn)
% at Tsinghua University, Beijing, China.
% check the parameters
if (nargin<1)
error('You must provide an image.');
elseif (nargin==1)
sw=0;
elseif (sw~=0 && sw~=1)
error('sw must be 0 or 1.');
end
data=reshape(IM,[],1);
[center,member]=fcm(data,3);
[center,cidx]=sort(center);
member=member';
member=member(:,cidx);
[maxmember,label]=max(member,[],2);
if sw==0
level=(max(data(label==1))+min(data(label==2)))/2;
else
level=(max(data(label==2))+min(data(label==3)))/2;
end
bw=im2bw(IM,level);
%testfcmthresh.m
clear;clc;
im=imread('mliver3.jpg');
fim=mat2gray(im);
level=graythresh(fim);
bwfim=im2bw(fim,0.1);
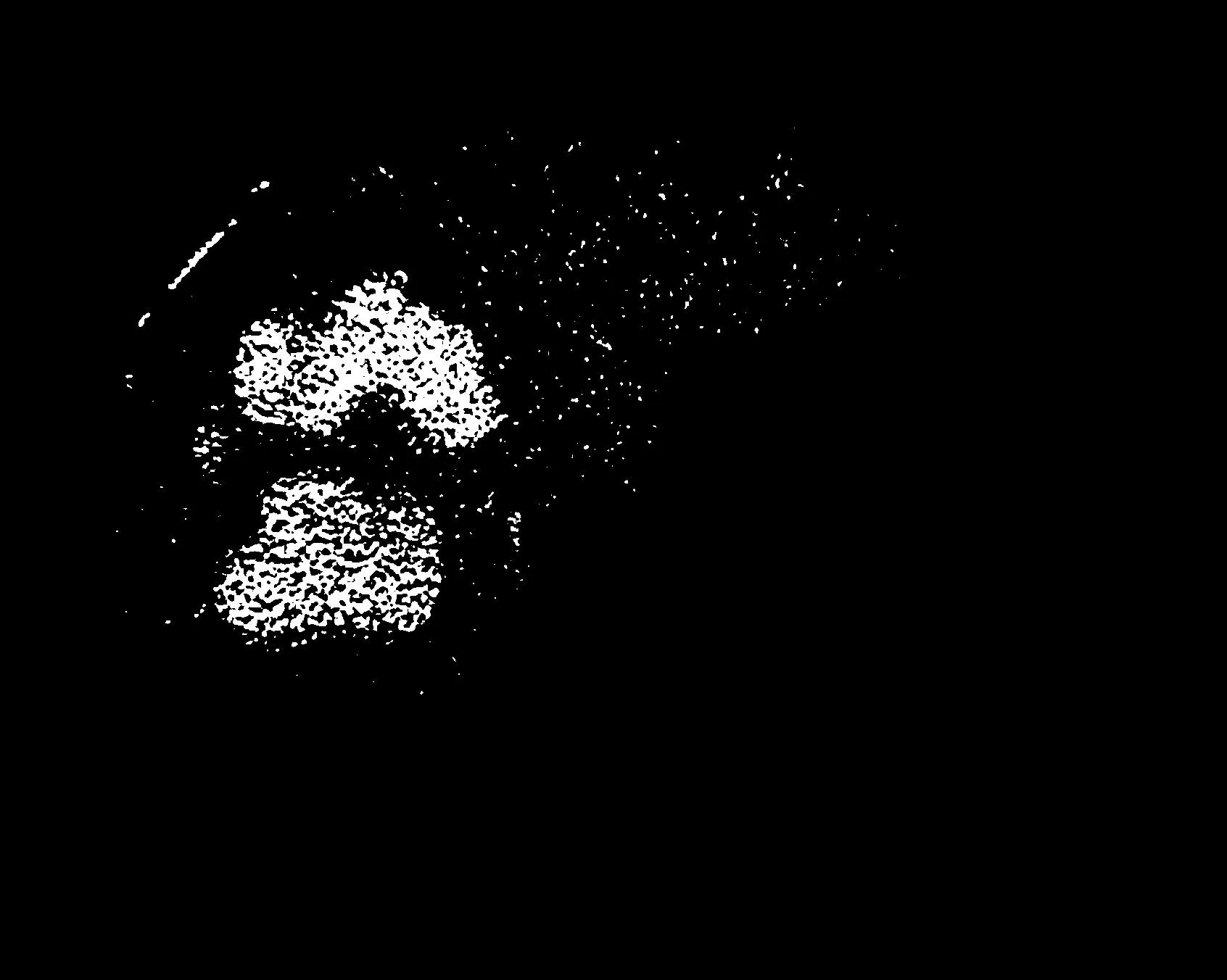
[bwfim0,level0]=fcmthresh(fim,0);
[bwfim1,level1]=fcmthresh(fim,1);
subplot(2,2,1);
imshow(fim);title('Original');
subplot(2,2,2);
imshow(bwfim);title(sprintf('Otsu,level=%f',level));
subplot(2,2,3);
imshow(bwfim0);title(sprintf('FCM0,level=%f',level0));
subplot(2,2,4);
imshow(bwfim1);title(sprintf('FCM1,level=%f',level1));
% imwrite(bwfim1,'fliver6.jpg');