- 如果每个时间步流过非零梯度,则每个时间步都对学习有贡献 - 即,由于考虑了每个输入时间步,所以导致的梯度源于整个序列影响权重更新。 - 如上所述,RNN 不再忽略长序列的部分,并被迫从中学习。
...但是我该如何在Keras/TensorFlow中实际可视化这些梯度呢?一些相关答案朝着正确方向,但它们似乎无法处理双向RNN,并且仅展示如何获得层的梯度,而不是如何有意义地可视化它们(输出是一个三维张量 - 我该如何绘制它?)
梯度可以相对于权重或输出获取 - 我们需要后者。此外,为了获得最佳结果,需要进行特定于架构的处理。下面的代码和解释涵盖了Keras/TF RNN的每种可能情况,并且应该可以轻松扩展到任何未来的API更改。
完整性: 显示的代码是简化版本 - 完整版本可在我的存储库See RNN中找到(本文包含更大的图像);包括以下内容:
from keras和from tf.kerasI/O维度 (所有RNNs):
(batch_size, timesteps, channels) - 或者等效的(samples, timesteps, features)channels/features现在是# of RNN units,和:return_sequences=True --> timesteps_out = timesteps_in (为每个输入时间步输出一个预测)return_sequences=False --> timesteps_out = 1 (仅在处理的最后一个时间步输出预测)可视化方法:
# for below examples
grads = get_rnn_gradients(model, x, y, layer_idx=1) # return_sequences=True
grads = get_rnn_gradients(model, x, y, layer_idx=2) # return_sequences=False
例子 1:一个样本,使用单向LSTM模型,6个神经元 -- return_sequences=True,训练了20个迭代周期。
show_features_1D(grads[0], n_rows=2)
例子 2: 所有 (16) 样本,单向 LSTM,6 个单位 -- return_sequences=True,训练了 20 次迭代
show_features_1D(grads, n_rows=2)
show_features_2D(grads, n_rows=4, norm=(-.01, .01))
例子 3: 所有 (16) 样本,单向 LSTM,6 个单位 -- return_sequences=True,训练了 200 次迭代
show_features_1D(grads, n_rows=2)
show_features_2D(grads, n_rows=4, norm=(-.01, .01))
例子 4: 2D vs. 1D,单向 LSTM: 256 个单位,return_sequences=True,训练了 200 次迭代
show_features_1D(grads[0])
show_features_2D(grads[:, :, 0], norm=(-.0001, .0001))
例5:双向GRU,256个单元(总共512个) - return_sequences=True,训练400次。
show_features_2D(grads[0], norm=(-.0001, .0001), reflect_half=True)
norm,因为近似相同的损失派生梯度被分布在更多参数上(因此平方数值平均值较小)。
例6:0D,所有(16)样本,uni-LSTM,6个单元 -- return_sequences=False,训练200次
show_features_0D(grads)
return_sequences=False仅利用最后一个时间步的梯度(仍源自所有时间步,除非使用截断的BPTT),需要一种新方法。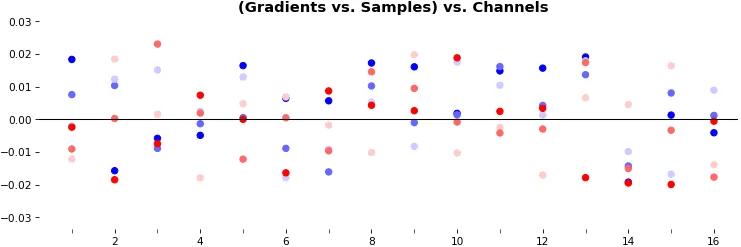
例7:LSTM vs. GRU vs. SimpleRNN,unidir,256个单元 -- return_sequences=True,训练250次
show_features_2D(grads, n_rows=8, norm=(-.0001, .0001), show_xy_ticks=[0,0], show_title=False)
可视化函数:
def get_rnn_gradients(model, input_data, labels, layer_idx=None, layer_name=None,
sample_weights=None):
if layer is None:
layer = _get_layer(model, layer_idx, layer_name)
grads_fn = _make_grads_fn(model, layer, mode)
sample_weights = sample_weights or np.ones(len(input_data))
grads = grads_fn([input_data, sample_weights, labels, 1])
while type(grads) == list:
grads = grads[0]
return grads
def _make_grads_fn(model, layer):
grads = model.optimizer.get_gradients(model.total_loss, layer.output)
return K.function(inputs=[model.inputs[0], model.sample_weights[0],
model._feed_targets[0], K.learning_phase()], outputs=grads)
def _get_layer(model, layer_idx=None, layer_name=None):
if layer_idx is not None:
return model.layers[layer_idx]
layer = [layer for layer in model.layers if layer_name in layer.name]
if len(layer) > 1:
print("WARNING: multiple matching layer names found; "
+ "picking earliest")
return layer[0]
def show_features_1D(data, n_rows=None, label_channels=True,
equate_axes=True, max_timesteps=None, color=None,
show_title=True, show_borders=True, show_xy_ticks=[1,1],
title_fontsize=14, channel_axis=-1,
scale_width=1, scale_height=1, dpi=76):
def _get_title(data, show_title):
if len(data.shape)==3:
return "((Gradients vs. Timesteps) vs. Samples) vs. Channels"
else:
return "((Gradients vs. Timesteps) vs. Channels"
def _get_feature_outputs(data, subplot_idx):
if len(data.shape)==3:
feature_outputs = []
for entry in data:
feature_outputs.append(entry[:, subplot_idx-1][:max_timesteps])
return feature_outputs
else:
return [data[:, subplot_idx-1][:max_timesteps]]
if len(data.shape)!=2 and len(data.shape)!=3:
raise Exception("`data` must be 2D or 3D")
if len(data.shape)==3:
n_features = data[0].shape[channel_axis]
else:
n_features = data.shape[channel_axis]
n_cols = int(n_features / n_rows)
if color is None:
n_colors = len(data) if len(data.shape)==3 else 1
color = [None] * n_colors
fig, axes = plt.subplots(n_rows, n_cols, sharey=equate_axes, dpi=dpi)
axes = np.asarray(axes)
if show_title:
title = _get_title(data, show_title)
plt.suptitle(title, weight='bold', fontsize=title_fontsize)
fig.set_size_inches(12*scale_width, 8*scale_height)
for ax_idx, ax in enumerate(axes.flat):
feature_outputs = _get_feature_outputs(data, ax_idx)
for idx, feature_output in enumerate(feature_outputs):
ax.plot(feature_output, color=color[idx])
ax.axis(xmin=0, xmax=len(feature_outputs[0]))
if not show_xy_ticks[0]:
ax.set_xticks([])
if not show_xy_ticks[1]:
ax.set_yticks([])
if label_channels:
ax.annotate(str(ax_idx), weight='bold',
color='g', xycoords='axes fraction',
fontsize=16, xy=(.03, .9))
if not show_borders:
ax.set_frame_on(False)
if equate_axes:
y_new = []
for row_axis in axes:
y_new += [np.max(np.abs([col_axis.get_ylim() for
col_axis in row_axis]))]
y_new = np.max(y_new)
for row_axis in axes:
[col_axis.set_ylim(-y_new, y_new) for col_axis in row_axis]
plt.show()
def show_features_2D(data, n_rows=None, norm=None, cmap='bwr', reflect_half=False,
timesteps_xaxis=True, max_timesteps=None, show_title=True,
show_colorbar=False, show_borders=True,
title_fontsize=14, show_xy_ticks=[1,1],
scale_width=1, scale_height=1, dpi=76):
def _get_title(data, show_title, timesteps_xaxis, vmin, vmax):
if timesteps_xaxis:
context_order = "(Channels vs. %s)" % "Timesteps"
if len(data.shape)==3:
extra_dim = ") vs. Samples"
context_order = "(" + context_order
return "{} vs. {}{} -- norm=({}, {})".format(context_order, "Timesteps",
extra_dim, vmin, vmax)
vmin, vmax = norm or (None, None)
n_samples = len(data) if len(data.shape)==3 else 1
n_cols = int(n_samples / n_rows)
fig, axes = plt.subplots(n_rows, n_cols, dpi=dpi)
axes = np.asarray(axes)
if show_title:
title = _get_title(data, show_title, timesteps_xaxis, vmin, vmax)
plt.suptitle(title, weight='bold', fontsize=title_fontsize)
for ax_idx, ax in enumerate(axes.flat):
img = ax.imshow(data[ax_idx], cmap=cmap, vmin=vmin, vmax=vmax)
if not show_xy_ticks[0]:
ax.set_xticks([])
if not show_xy_ticks[1]:
ax.set_yticks([])
ax.axis('tight')
if not show_borders:
ax.set_frame_on(False)
if show_colorbar:
fig.colorbar(img, ax=axes.ravel().tolist())
plt.gcf().set_size_inches(8*scale_width, 8*scale_height)
plt.show()
def show_features_0D(data, marker='o', cmap='bwr', color=None,
show_y_zero=True, show_borders=False, show_title=True,
title_fontsize=14, markersize=15, markerwidth=2,
channel_axis=-1, scale_width=1, scale_height=1):
if color is None:
cmap = cm.get_cmap(cmap)
cmap_grad = np.linspace(0, 256, len(data[0])).astype('int32')
color = cmap(cmap_grad)
color = np.vstack([color] * data.shape[0])
x = np.ones(data.shape) * np.expand_dims(np.arange(1, len(data) + 1), -1)
if show_y_zero:
plt.axhline(0, color='k', linewidth=1)
plt.scatter(x.flatten(), data.flatten(), marker=marker,
s=markersize, linewidth=markerwidth, color=color)
plt.gca().set_xticks(np.arange(1, len(data) + 1), minor=True)
plt.gca().tick_params(which='minor', length=4)
if show_title:
plt.title("(Gradients vs. Samples) vs. Channels",
weight='bold', fontsize=title_fontsize)
if not show_borders:
plt.box(None)
plt.gcf().set_size_inches(12*scale_width, 4*scale_height)
plt.show()
完整的最简示例:请参阅存储库的README
额外的代码:
rnn_cell = model.layers[1].cell # unidirectional
rnn_cell = model.layers[1].forward_layer # bidirectional; also `backward_layer`
print(rnn_cell.__dict__)
为了更方便的代码,请查看存储库的rnn_summary
奖励知识点:如果在GRU上运行上述代码,您可能会注意到bias没有门;为什么呢?从文档中得知:
有两个变体。 默认值基于1406.1078v3,并在矩阵乘法之前将重置门应用于隐藏状态。另一个基于原始的1406.1078v1,顺序相反。
第二个变体与CuDNNGRU(仅GPU)兼容,并允许在CPU上进行推断。因此,它具有用于内核和recurrent_kernel的单独偏差。使用'reset_after'=True和recurrent_activation ='sigmoid'。