3个回答
12
一些时间:
set.seed(1001)
d <- sample(1:100000, 100000, replace=T)
d <- c(d, sample(d, 20000, replace=T)) # ensure many duplicates
mb <- microbenchmark::microbenchmark(
d[!(duplicated(d) | duplicated(d, fromLast=TRUE))],
setdiff(d, d[duplicated(d)]),
{tmp <- rle(sort(d)); tmp$values[tmp$lengths == 1]},
as.integer(names(table(d)[table(d)==1])),
d[!(duplicated.default(d) | duplicated.default(d, fromLast=TRUE))],
d[!(d %in% d[duplicated(d)])],
{ ud = unique(d); ud[tabulate(match(d, ud)) == 1L] },
d[!(.Internal(duplicated(d, F, F, NA)) | .Internal(duplicated(d, F, T, NA)))]
)
summary(mb)[, c(1, 4)] # in milliseconds
# expr mean
#1 d[!(duplicated(d) | duplicated(d, fromLast = TRUE))] 18.34692
#2 setdiff(d, d[duplicated(d)]) 24.84984
#3 { tmp <- rle(sort(d)) tmp$values[tmp$lengths == 1] } 9.53831
#4 as.integer(names(table(d)[table(d) == 1])) 255.76300
#5 d[!(duplicated.default(d) | duplicated.default(d, fromLast = TRUE))] 18.35360
#6 d[!(d %in% d[duplicated(d)])] 24.01009
#7 { ud = unique(d) ud[tabulate(match(d, ud)) == 1L] } 32.10166
#8 d[!(.Internal(duplicated(d, F, F, NA)) | .Internal(duplicated(d, F, T, NA)))] 18.33475
鉴于评论,请让我们看看它们是否都正确?
results <- list(d[!(duplicated(d) | duplicated(d, fromLast=TRUE))],
setdiff(d, d[duplicated(d)]),
{tmp <- rle(sort(d)); tmp$values[tmp$lengths == 1]},
as.integer(names(table(d)[table(d)==1])),
d[!(duplicated.default(d) | duplicated.default(d, fromLast=TRUE))],
d[!(d %in% d[duplicated(d)])],
{ ud = unique(d); ud[tabulate(match(d, ud)) == 1L] },
d[!(.Internal(duplicated(d, F, F, NA)) | .Internal(duplicated(d, F, T, NA)))])
all(sapply(ls, all.equal, c(3, 5, 6)))
# TRUE
- Raad
6
4不妨全力以赴
d[!(.Internal(duplicated(d, FALSE, FALSE, NA)) | .Internal(duplicated(d, FALSE, TRUE, NA)))] - rawr2最好先检查正确性(与测试用例的一致性),然后是鲁棒性(包括字符、整数、数字向量,从更新的问题中获得;names(tables(...)) 的解决方案会失败),最后再考虑性能。 - Martin Morgan
哎呀呀,第二名! - Señor O
2在更大的向量上进行基准测试会不会更加干净呢?另外,
do.call(all.equal, results)仅比较“results”的前两个元素;其他元素将作为其他参数传递。 - alexis_laz看起来我的解决方案最终是第一个。 - Bulat
这实际上是误导性的,至少使用10^6作为样本,由于这是一个单向量,我甚至会选择10^8。结果差异太大了! - Bulat
7
您可以使用
找到向量中相同值的数量是个好主意。
这里有很多替代方法:在向量中计算具有x值的元素数量 编辑 看完基准测试后,@NBATrends让我有些怀疑。理论上说,通过单次遍历计算项目数必须比原始的
我尝试使用
以下是三种解决方案在不同样本大小下的基准测试结果(仅列出了快速独特方法):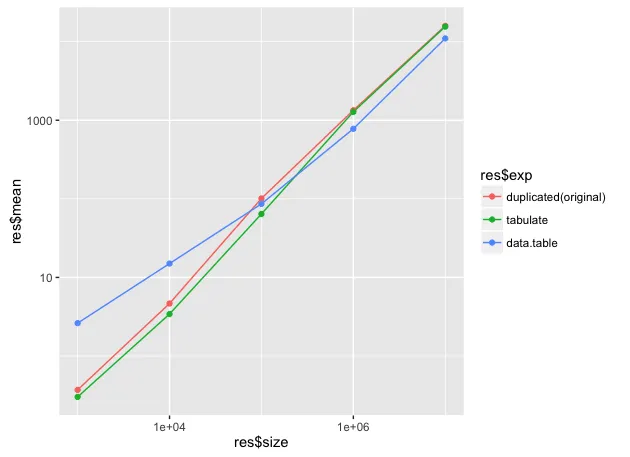 。可以看到,随着样本量的增长,data.table 开始表现出优异的性能(2倍)。以下是代码以进行复制:
。可以看到,随着样本量的增长,data.table 开始表现出优异的性能(2倍)。以下是代码以进行复制:
rle 函数来实现此操作:tmp <- rle(sort(d))
res <- tmp$values[tmp$lengths == 1]
找到向量中相同值的数量是个好主意。
这里有很多替代方法:在向量中计算具有x值的元素数量 编辑 看完基准测试后,@NBATrends让我有些怀疑。理论上说,通过单次遍历计算项目数必须比原始的
duplicated逻辑快约2倍。我尝试使用
data.table来实现这一点:library(data.table)
dt <- data.table(d)
res <- dt[, count:= .N, by = d][count == 1]$d
以下是三种解决方案在不同样本大小下的基准测试结果(仅列出了快速独特方法):
 。可以看到,随着样本量的增长,data.table 开始表现出优异的性能(2倍)。以下是代码以进行复制:
。可以看到,随着样本量的增长,data.table 开始表现出优异的性能(2倍)。以下是代码以进行复制:set.seed(1001)
N <- c(3, 4, 5, 6 ,7)
n <- 10^N
res <- lapply(n, function(x) {
d <- sample(1:x/10, 5 * x, replace=T)
d <- c(d, sample(d, x, replace=T)) # ensure many duplicates
dt <- data.table(d)
mb <- microbenchmark::microbenchmark(
"duplicated(original)" = d[!(duplicated(d) | duplicated(d, fromLast=TRUE))],
"tabulate" = { ud = unique(d); ud[tabulate(match(d, ud)) == 1L] },
"data.table" = dt[, count:= .N, by = d][count == 1]$d,
times = 1,unit = "ms")
sm <- summary(mb)[, c(1, 4, 8)]
sm$size = x
return(sm)
})
res <- do.call("rbind", res)
require(ggplot2)
##The values Year, Value, School_ID are
##inherited by the geoms
ggplot(res, aes(x = res$size, y = res$mean, colour=res$exp)) +
geom_line() + scale_x_log10() + scale_y_log10() +
geom_point()
- Bulat
4
1就此而言,标准的
data.table 语法/做法是 dt[, if(.N == 1) .SD, by = d]。 - eddi从语法角度来看,它很有趣,但在速度方面并不高效。 - Bulat
@Bulat,哪个更快很大程度上取决于重复项的分布。 - eddi
@eddi,我想不出为什么会这样。 - Bulat
0
你可以使用一个集合操作:
d <- c(1,2,3,4,1,5,6,4,2,1)
duplicates = d[duplicated(d)]
setdiff(d, duplicates)
[1] 3 5 6
(不确定这是否比上面的代码更有效,但从概念上看似乎更清晰)
- djq
网页内容由stack overflow 提供, 点击上面的可以查看英文原文,
原文链接
原文链接
d[!(d %in% d[duplicated(d)])]。 - lmoduplicated.default可以节省约50%的时间。 - Señor O{ ud = unique(d); ud[tabulate(match(d, ud)) == 1L] }。 - Martin Morgan